- Details
- Written by Administrator
- Category: molecular biology
- Hits: 3983
1 - 9
4D Nucleome project. The goal of this program is to study the three-dimensional organization of the nucleus in space and time (the 4th dimension).
1000 Genomes Project created a catalogue of common human genetic variation, using openly consented samples from people who declared themselves to be healthy. The reference data resources generated by the project remain heavily used by the biomedical science community. The International Genome Sample Resource (IGSR) maintains and shares and updates the human genetic variation resources built by the 1000 Genomes Project.
100,000 genomes project. a UK Government project that is sequencing whole genomes from National Health Service patients. The project is focusing on rare diseases, some common types of cancer, and infectious diseases.
A - E
Alpha-keto acid. Keto acids that have the keto group adjacent to the carboxylic acid.
| Alpha-keto acid | No C | Transamination / Enzyme | Reduction/ Enzyme | |
| Pyruvic acid | 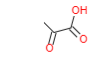 |
3 |
Alanine + Alpha-keto glutarate <----> Pyruvic acid + Glutamate Alanine aminotransferase (ALAT) |
Pyruvic acid + NADH +
H+ <----> Lactate + NAD+ Lactate dehydrogenase |
| Oxaloacetic acid | 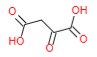 |
4 |
Aspartate + Alpha-keto glutarate <----> Oxaloacetic acid + Glutamate Aspartate aminotransferase (ASAT) |
Oxaloacetic acid + NADH + H+ <----> Malate + NAD+ Malate dehydrogenase |
| Alpha-keto glutarate | 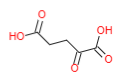 |
5 |
Amino acid + Alpha-keto glutarate <----> Alpha-keto acid + Glutamate |
Alpha-keto glutarate + CO2
+ NADH + H+ <----> Isocitrate + NAD+ Isocitrate dehydrogenase |
Amino acid, proteinogenic. L-stereo isomer alpha-amino acid that is incorporated into proteins during translation.
2nd column: three letter abbrevation amino acid
3rd column: one letter abbrevation amino acid
4th column: E = essential amino acid, C = conditional essential, blank = not essential
Hydrophobic sidechain, aliphatic
|
Alanine |
Ala | A | 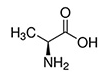 |
Abundant and versatile, it is more stiff than glycine, but small enough to pose only small steric limits for the protein conformation. |
|
| Isoleucine | Ile | I | E | 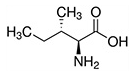 |
Isoleucine, leucine, and valine have large aliphatic hydrophobic side chains. Their molecules are rigid, and their mutual hydrophobic interactions are important for the correct folding of proteins, as these chains tend to be located inside of the protein molecule. |
| Leucine | Leu | L | E | 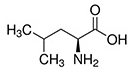 |
|
| Valine | Val | V | E | 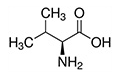 |
|
| Methionine | Met | M | E | 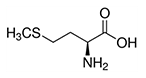 |
- Always the first amino acid to be incorporated into a protein, it is sometimes removed after translation. - Like cysteine, it contains sulfur, but with a methyl group instead of hydrogen. This methyl group can be activated, and is used in many reactions where a new carbon atom is being added to another molecule (S-Adenosylmethionine, SAM-e). |
Hydrophobic sidechain, aromatic. Like isoleucine, leucine, and valine, these are large hydrophobic amino acids and tend to orient towards the interior of the folded protein molecule.
| Phenylalanine | Phe | F | E | 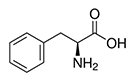 |
- can be converted to tyrosine - phenylketonuria (PKU) is an inborn error of metabolism caused by genetic low levels of phenylalanine hydroxylase resulting in toxic levels of phenylalanine, leading to retarded brain development. |
| Tryptophan | Trp | W | E | 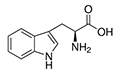 |
precursor of serotonine precursor of nicotinamide adenine dinucleotide (NAD) |
| Tyrosine | Tyr | Y | C | 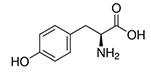 |
precursor of melanin, epinephrine and thyroid hormones |
Polar uncharged sidechain
| Asparagine | Asn | N | 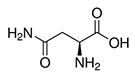 |
- Both contain an amide group (RC(=O)NR'R'', in this case R' and R'' are hydrogen atoms) - Glutamine is the most abundant free AA in plasma (> 20 % of free AA) |
|
| Glutamine | Gln | Q | 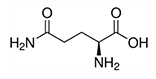 |
||
| Serine | Ser | S | 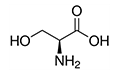 |
- the hydrogen of their hydroxyl groups is easy to remove so serine and threonine often act as hydrogen donors in enzymes - they are very hydrophilic, so they are abundant in outer regions of soluble proteins |
|
| Threonine | Thr | T | E | 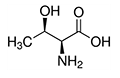 |
Positive charged sidechain, basic
| Arginine | Arg | R | C | 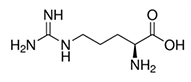 |
abundant in histone |
| Histidine | His | H | E | 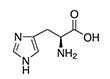 |
|
| Lysine | Lys | K | E | 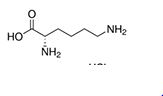 |
abundant in histone |
Negative charged sidechain, acidic
| Aspartic acid | Asp | D | 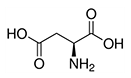 |
- these are usually located on the outer surface of a protein, making it water-soluble - glutamic acid has a longer, slightly more flexible side chain - aspartate is the ionic form of aspartic acid - aspartate is part of the malate-aspartate shuttle - glutamate is the ionic form of glutamic acid - glutamate is the most abundant excitatory neurotransmitter |
|
| Glutamic acid | Glu | E | C | 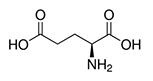 |
Special cases
| Cysteine | Cys | C | C | 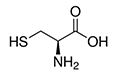 |
- the sulphur atom bonds readily to heavy metal ions - two cysteins can join into a disulfide bond, which has a stabilizing effect on the tertiary structure of proteines. Cysteine is therefore often found in proteines: * that have to function in harsh environments (e.g. digestive enzymes) * with a structural function (e.g. keratin) * that are too small to hold a stable shape on their own (e.g. insulin) |
| Selenocysteine | Sec | U | 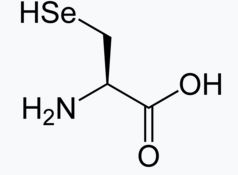 |
||
| Glycine | Gly | G | C | 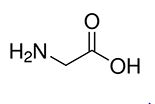 |
|
| Proline | Pro | P | 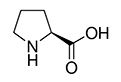 |
Anaplerosis. The process of replenishing intermediates of the citric acid cycle. Beside being the central process in energy metabolism, the citric acid cycle also functions in biosynthetic pathways in which intermediates leave the cycle to be converted primarily to glucose, fatty acids, or non-essential amino acids. The removed citric acid intermediates must be replaced to permit continued function (anaplerosis). Conversely, 4- and 5-carbon intermediates enter the citric acid cycle during the catabolism of amino acids. Because the citric acid cycle cannot fully oxidize 4- and 5-carbon compounds (cannot act as a carbon sink) intermediates must be removed from the cycle, a process called cataplerosis. A balance between the reciprocal processes anaplerosis and cataplerosis is thus necessary (Owen 2002).
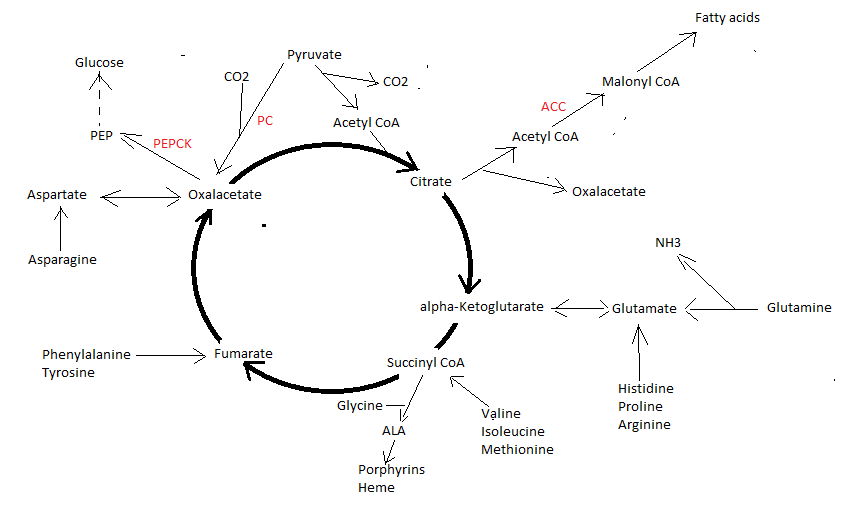
Citric acid cycle with some major anaplerotic and cataplerotic reactions. Anaplerosis includes net entry of amino acids into the cycle and the generation of oxaloacetate from pyruvate via the "archetypical anaplerotic enzyme" pyruvate carboxylase (PC). Cataplerotic reactions include conversion of oxaloacetate to PEP by the "cataplerotic enzyme" PEPCK (linkage to gluconeogenesis), linkage to lipogenesis (ACC = Acetyl-CoA carboxylase) and conversion of succinyl CoA to ALA (Aminolevulinic acid) leading tot porphyrin and heme synthesis.
Assembly. A database providing information on the structure of assembled genomes, assembly names and other meta-data, statistical reports, and links to genomic sequence data.
Bioinformatics, centres of excellence.
see European Molecular Biology Laboratory - European Bioinformatics Intitute (EMBL-EBI)
see National Centre for Biotechnology Information (NCBI)
see National Institute of Genetics (NIG)
Biotin (vitamin B7, formerly vitamin H)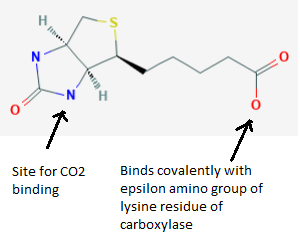
Biological functions:
- Acts as a carrier of carbon dioxide, as a prosthetic group in enzymes involved in biotin-dependent carboxylation. One side is covalently bound to the enzyme, by the enzyme holocarboxylase synthetase (HLCS), forming a long arm that moves the intermediate (CO2) from one active site to the next.
- Histone biotinylation: are a side effect of HLCS being in close proximity to histones and play no direct role in gene repression (Mock 2017).
- Biotinylation of other proteins: is probably more prevalent than previously thought. Especially proteins from the heat-shock protein (HSP) superfamily were found to be biotinylated.
Deficiency
Frank symtomatic deficiency is rare, because biotin is contained in so many foods. The following conditions are related with deficiency:
- Pregnancy may often lead to maginal biotin deficiency, which probably can have a teratogenic effect.
- Consuming large quantities of uncooked egg white, which is known to contain the protein avidin, which strongly binds biotin. The strong binding of biotin and avidin is in fact used in many molecular biologic techniques, where avidin is mostly exchanged for the bacterial streptavidin.
- Metabolic disorders: for instance biotinidase deficiency, holocarboxylase synthetase deficiency
Therapeutic uses
High dose biotin therapy is used in biotin-thiamin-responsive basal ganglia disease (BTBGD). It may also be an option in slowing down advancement of progressive mutiple sclerosis.
Other issues
- Biotin supplement is often promoted as unsupported claims for strenghtening hair and nails.
- Using high dose dietary supplements of biotin can give clinically significant interference with diagnostic blood tests that use biotin-streptavidin technology (e.g. thyroid hormones). Depending on the test, false results can be too high or to low. Sometimes combined lab results seem falsely to support each other (e.g. high FT4 with low TSH).
Basic Local Alignment Search Tool (BLAST). A suit of programs provided by NCBI for aligning query sequences against those present in a selected target database. BLAST for beginners
Carboxylation chemical reaction in which a carboxylic acid (R-(C(=O)OH)) is produced by treating a substrate with carbon dioxide
Biotin-dependent carboxylation:
| Enzyme | abbr | Reaction | Pathway |
| Acetyl-CoA carboxylase | ACC | acetyl-CoA -> malonyl-CoA | fatty acid synthesis |
| Methylcrotonyl CoA carboxylase | MCC | leucine metabolism | |
| Propionyl-CoA carboxylase | PCC | propionyl-CoA -> (S)-methylmalonyl-CoA |
succinyl-CoA production |
| Pyruvate carboxylase | PC | pyruvate -> oxaloacetate |
Biotin-independent carboxylation:
1. Gamma carboxylation. Vitamin K dependent posttranslational modification of glutamate to γ-carboxyglutamate. This reaction is necessary for some proteins of the coagulation pathway (factors II, VII, IX, and X, protein C, protein S and protein Z).
2. Formation of carbamoyl phosphate in the urea cycle by carbamoyl phosphate synthetase I (CPS I).
3. Introduction of carbon dioxide in 6th step of purine synthesis .
4. Conversion of pyruvate to malate by malate dehydrogenase.
Cataplerosis. See Anaplerosis
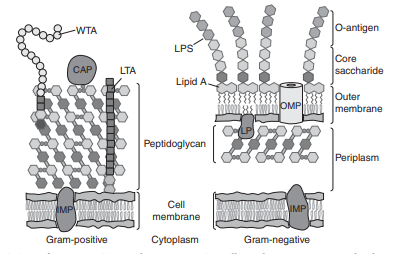
Cell envelope complex multilayered structure that encloses the cytoplasm of bacteria. Using Gram staining two mayor types of bacterial cell envelopes can be distinguished: Gram-positive and Gram-negative. The Gram-positive cell envelope comprises the (inner) cell membrane and a cell wall made of a thick layer of peptidoglycan. The Gram-negative cell envelope has an inner cell membrane, a thin layered peptidoglycan cell wall and an outer cell membrane:
Depiction of Gram-positive and Gram-negative cell envelopes: CAP = covalently attached protein; IMP, integral menbrane proteine; LP, lipoprotein; LPS, lipopolysaccharide; LTA, lipoteichoic acid; OMP, outer membrane protein; WTA, wall teichoic acid (Silhavy 2010)
Cell signaling ability of cells to receive, process, and transmit signals with its environment and with itself. Signals can be physical or chemical. Chemical signaling can be classified as:
1. Autocrine. The chemical signal secreted by a cell bind to receptors on that same cell.
2. Juxtacrine or contact-dependent signaling. Takes place by direct contact between two cells.
3. Intracrine. The chemical signal acts inside a cell.
4. Paracrine. The produced chemical signal acts on receptors of nearby cells.
5. Endocrine. The produced chemical signal acts on distant cells after transport by the circulatory system.
In general three components can be recognised in cell signaling:
1. Physical or chemical activation of a receptor
2. Signal transduction: activation of the receptor triggers a change in the activity or state of a cell
3. Response by regulation of a downstream cellular process, e.g. regulation of transcription or regulation of a metabolic process.
Cellular respiration. Process by which organic nutrients (most common: glucose, amino acids, fatty acids) are oxidised in the presence of an electron acceptor to produce energy, which is stored as ATP.
Cellulair respiration that uses molecular oxygen (O2) as electron acceptor is called aerobic respiration. Anearobic respiration uses other electron acceptors, for example nitrate, iron, fumarate, sulphate or elemental sulphur. These have smaller reduction potentials than O2, so anaerobic respiration is less efficient than aerobic respiration.
Both aerobic and anaerobic respiration use highly reduced chemical compounds like NADH or FADH2 to establish an electrochemical gradient (often a proton gradient) across a membrane. The reduced chemical compounds are oxidised by series of respiratory integral membrane proteins with sequentially increasing reduction potentials, with the final electron acceptor oxygen or another chemical.
Anaerobic respiration has to be differentiated from fermentation, which does not use an electrochemical gradient.
Cereal (graan) Any grass cultivated for its edible grain (korrel), which is composed of an endosprem (kiemwit), a bran (zemelen) and a germ (graankiem). Cereals are the most important crops in the world (Shewry 2002). The seven most important cereals are (data from FAOSTAT):
| Cereal |
Worldwide production 2021 (millions of metric tons) |
Notes |
| Maize or corn (mais) | 1210 | A staple food of people in the Americas, Africa, and of livestock worldwide. A large portion of maize crops are grown for purposes other than human consumption. |
| Rice (rijst) | 787 | The primary cereal of tropical and some temperate regions. |
| Wheat (tarwe) | 770 | The primary cereal of temperate regions. It has a worldwide consumption but it is a staple food of North America, Europe, Australia, New Zealand, Argentina, Brazil and much of the Greater Middle East. |
| Barley (gerst) | 145 | Grown for malting and livestock on land too poor or too cold for wheat. |
| Millet (gierst) | 30 | A group of similar but distinct cereals that form an important staple food in Asia and Africa. |
| Oats (haver) | 22 | Popular worldwide as a breakfast food and livestock feed. |
| Rye (rogge) | 13 | Important in cold climates. Rye grain is used for flour, bread, beer, some whiskeys, some vodkas, and animal fodder. |
Chromosome conformation capture. A family of biochemical techniques (C-technologies) to determine the physical interaction of genome regions (Hakim 2012)
Cis-regulatory element (CRE), opposite of Trans-regulatory element (TRE). Region of non-coding DNA that regulates the transcription of a nearby gene. CREs are positioned on the same molecule ( cis = on this side) as the regulated gene, often, but not always upstream.
Citric acid cycle (tricarboxylic acid (TCA) cycle, Krebs cycle).
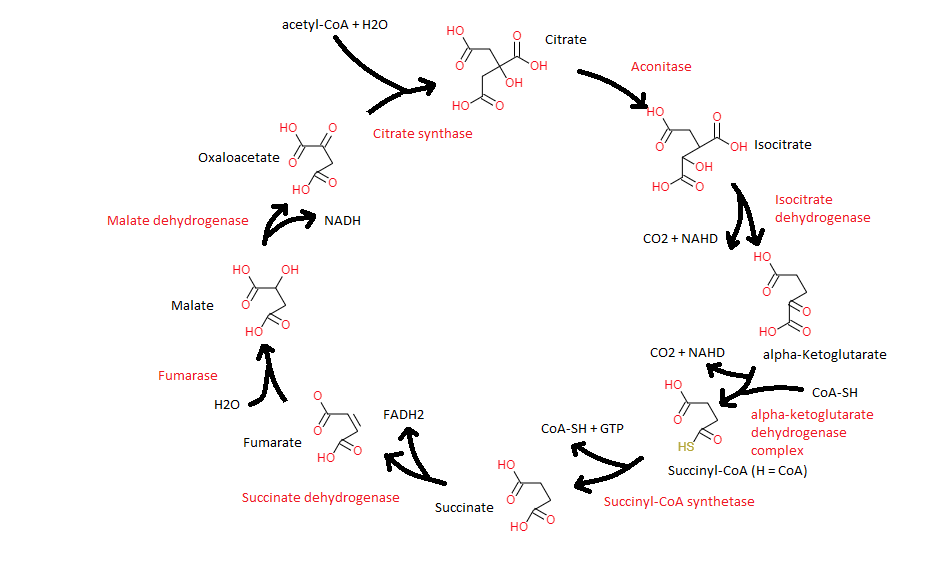
1. Energy production. One of the major parts of cellular respiration, where an acetylgroup, in the form of acetyl-CoA, is oxidized into two carbon dioxide molecules, conserving the released energy in the reduced electron cariers NADH and FADH2. The chemical rationale behind the citric acid cycle is that plain decarboxylation of the acetyl group would yield carbon dioxide and methane (CH4). Methane being rather stable, most organisms lack the cofactors and enzymes to oxidize it. Methylene groups (-CH2-) are much easier to metabolize, especially when adjacent to a carbonyl group (R-(C=O)-R)), because the of relative positive charge of the carbon of the carbonyl group, due to the electron withdrawing property of the oxygen of the carbonyl group (see alpha-keto acid).
2. Evolution - metabolic hub. In evolution anaerobic organisms preceded aerobic organisms. So probably in early anaerobic organisms, as seen in modern anaerobics, parts of the citric acid reactions where used to produce biosynthetic precursors. In aerobic organisms the citric acid cycle is an amphibolic pathway, serving both catabolic and anabolic processes (see anaplerosis ).
Cluster of Differentiation (CD). Antigenic cell surface molecules expressed on leukocytes and other cells relevant for the immune system. The nomenclature and definition of CDs is coordinated by the Human Cell Differentiation Molecules organisation.
Clusters of Orthologous Groups of proteins (COGs) phylogenetic classification of proteins encoded in complete genomes. List COG classes.
Structure
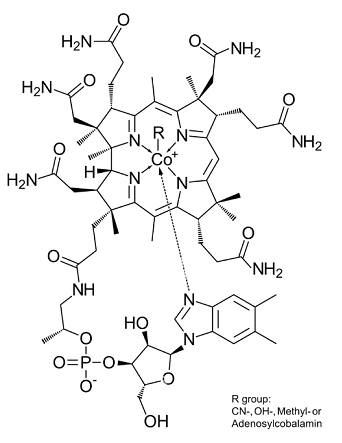
Cobalamin (Cbl) is nature's most chemically complex coenzyme:
1. Central cobalt atom, which can exist as Co(III) (can form 6 bonds), Co(II) (5 bonds) and Co(I) (4 bonds).
2. Four bonds of cobalt are always occupied by the nitrogen atoms of the planar corrin ring.
3. Additionally cobalt may binds a lower axial ligand, the dimethylbenzimidazole(DMB) moiety, which attaches back to the corrin ring.
4. The cobalt atom may also bind an upper axial ligand (R-group) e.g.:
- adenosyl- or methyl- (as cofactor)
- glutathionyl- or hydroxo- (other physiologically important)
- cyano- (common in pharmaceutical preparations)
Source and intake
Complete Cbl synthesis is limited to selected Eubacteria and Archaea, and requires more than 25 different steps at large energetic cost. For humans sources are limited to animal products (meat, shellfish, liver, poultry, eggs, dairy products). Approximately 20 human genes are known to be involved in absorption, selection, transport, modification, and utilization of Cbl acquired from the diet. In the stomach Cbl is freed by proteolytic cleavage of the food mass, and is bound by the protein haptocorrin, which is thought to protect Cbl from the acidic environment. Haptocorrin is degraded in the duodenum by proteases from the pancreas, after which Intrinsic Factor (IF), synthesized by parietal cells of the stomach, sequesters the released Cbl. (Froese et al 2019).
Codex Alimentarius or "Food Code" is a collection of standards, guidelines and codes of practice adopted by the Codex Alimentarius Commission. The Commission, also known as CAC, is the central part of the Joint FAO/WHO Food Standards Programme and was established by FAO and WHO to protect consumer health and promote fair practices in food trade.
CoGe. platform for performing Comparative Genomics research
Contig (from contiguous) is a set of overlapping DNA segments that together represent a consensus region of DNA.
CpG island (CGI). Region of clustered CpG: a cytosine nucleotide followed by a guanine nucleotide in the linear sequence of bases along its 5' → 3' direction. Cytosines in CpG dinucleotides can be methylated to form 5-methylcytosines. Enzymes that add a methyl group are called DNA methyltransferases. In mammals, 70% to 80% of CpG cytosines are methylated.CpG islands are often located at promoters.
Cytokine. Small molecule (glyco)proteins produced by a broad range of cells, notably but not exclusively immune cells, and are important in cell signaling.
Structural classification (using InterPro)
| Homologous Superfamily | Family | Subfamily |
| Interleukine- 2 | ||
|
Interleukine-6/GCSF/MGF |
Interleukine-6 | |
|
GCSF/MGF granulocyte colony-stimulating factor (GCSF) and myelomonocytic growth factor (MGF) |
||
| Interleukine - 10 | ||
|
Interferon alpha/beta/delta |
||
| Interferon gamma | ||
|
GM-CSF Granulocyte-macrophage colony-stimulating factor |
||
| Erythropoietin/thrombopoietin | Erythropoietin | |
| Thrombopoietin | ||
| Somatotropin/prolactine | Somatotropin | |
| Prolactine | ||
| Leptin | ||
| Cytokine IL-1/FGF | Interleukine-1 family | Interleukine-18 |
| Fibroblast growth factor family | ||
| Cystine-knot cytokine
|
Interleukin-17 family | |
| Transforming growth factor-beta | ||
| Nerve growth factor-like | Brain derived neurotrophic factor | |
| Tumour necrosis factor-like domain superfamily | Tumour necrosis factor alpha | |
|
|
Lymfotoxin-alpha (TNF-beta) | |
| Lymfotoxin-beta | ||
| Chemokine interleukine-8-like superfamily | C Chemokine | |
| CC Chemokine | ||
| CXC Chemokine | ||
|
|
CX3C Chemokine |
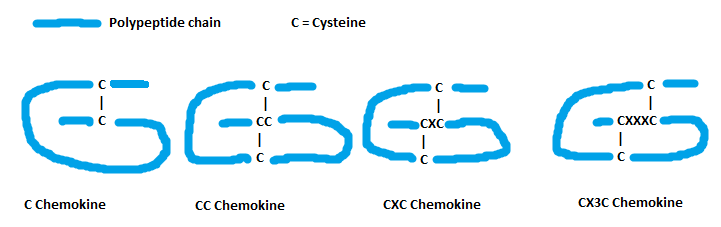
Functional classification
| 1. Pro-Inflammatory | |||
| Innate and Acute Phase Responses | |||
| Cytokine | Source | Major Target | Function |
| INF-α/β | Virally infected nucleated cells | Cells surrounding virally infected cells | Induce antiviral state, activate NK cells, enhance Cell Mediated Immunity |
| IL-1β | Macrophages, DCs, Fibroblasts, Epi/Endothetium | T and B cells/ PMNs/ CNS/ liver | Promote inflammatory/ acute phase response/ promote fever |
| TNF-α | Macrophages/ T cells/ NK cells | Acute phase response/ inflammation/ fever/ wasting (cachexia) | |
| IL-6 | idem + T and B cells | T and B cells/ hepatocytes | Acute phase reactants/ fever/ T and B cell growth |
| Inflammatory Innate to Adaptive Responses | |||
| INF-γ | CD4 Th1 cells, NK cells | Macrophages, DCs, B and T cells | Activate Macrophages for intracellular microbes/ promote IgG class switch/ promote Th1/ Inhibit Th2 |
| IL-17 | CD4 Th17 cells | Epi/endothelium/ Neutrophils/ Fibroblasts | Recruit Neutrophils for extracellular bacteria/fungi/ Promote inflammation |
|
TNF-β (Lymfotoxin-α) |
CD4 Th1 cells | PMNs, Tumor Cells | Kill tumors/ activate PMNs/ Activate Endothelium allowing trafficking |
| IL-2 | CD4 T cells (Th1, Th0) | Lymphocytes | Lymphocyte proliferation |
| IL-12/IL-23 | DCs, Macrophages | NK Cells, CD4 Th1/Th17 | Activate T cell INF-γ or IL-17 production |
| 2. Allergy and Helminth Infections - Th2 (Less inflammatory than Th1/Th17) | |||
| IL-4 | CD4 T cells (Th0, Th2, Tfh, ILCs) | B and T cells | B and T cell growth and differentiation/ IgG, IgA, IgE production/ Th2 response/ Allergic responses |
| IL-5 | CD4 Th2 | B cells and Eosinophils (for Helminth infections) | |
| IL-10 | CD4 Th2 and Treg | B cells, CD4 Th1/Th17 cells | B cell growth/ inhibit Th1 and Th17 responses |
| 3. Immunosuppressive/ Anti-inflammatory - T regulatory (Tregs) | |||
| IL-10 | Treg cells/ Th2 | B cells, CD4 Th1/Th17 cells | B cell growth/ inhibit Th1 and Th17 responses |
| TGF-β | CD4 Tregs | B and T cells, Macrophages, DCs | Immunosuppression of all immune responses/ promote oral tolerance/ wound healing |
| 4. Growth and Differentiation | |||
| GM-CSF & M-GSF | Stromal cells/ T cells | Bone marrow progenitor cells/ Stem cells/ Precursor cells | Hematopoiesis/ directed growth and differentiation of Monocytes and Granulocytes |
| IL-3 | CD4 T cells/ keritonocytes | ||
| IL-7 | Bone marrow, stroma | Growth of Pre-B cells, T cells and NK cells | |
| IL-2 | cd4 T cells | Lymphocytes | Proliferation |
| 5. Chemokines (Chemotactic cytokines) | |||
| IL-8/ CXCL8 | Fibroblasts/ Neutrophils/ Macrophages | Phagocytes | Recruit these cells to the site of inflammation |
| C5a | Complement cascade | ||
| IL-17 | CD4 Th17 | ||
| Over 30 others | Many cells | Neutrophils, B and T cells, Macrophages, DCs, NK cells, Mast cells etc | |
see Assembly
see Ensembl
see Genome aggregation database (gnomAD)
see Genome browsers
see Human genes and genetic diseases, databases
see Microbial Genome Database for Comparative Analysis
see Integrated Microbial Genomes and Microbiomes
see Model organism
see Pathguide
see Zoonomia
De novo gene birth is the process by which new genes evolve from non-coding DNA sequences.
DNA methylation. Methylation of nucleotides by DNA methyltransferases provide a molecular means to reversibly mark genomic DNA. Bacteria can methylate adenosine or cytosine. In eukaryotes DNA methylation only (mainly ?) occurs at cytosine. Methylation is not always essential to eukaryotic gene regulation as it is absent in many organisms (e.g. Caenorhabditis elegans). Vertebrates are unique in that cytosine methylation occurs throughout the entire genome. By contrast, in plants and invertebrates only specific genomic elements are targeted (repetitive DNA en actively transcribed sequences).
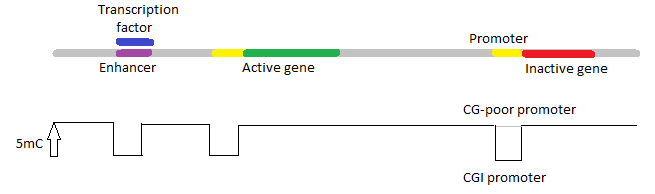
Genomic distribution of methylated cytosine in a typical invertebrate and vertebrate genome. The representative genomic region includes an example of an active and an inactive gene with proximal (promoter) and distal (enhancer) regulatory regions. The height of the bar indicates the relative proportion of DNA methylation (5-methylcytosine, 5mC) that is observed in each region. CpG islands (CGIs), which often overlap with promoter regions, generally remain unmethylated, whereas CG-poor promoters are methylated when not active (Schübeler 2015).
DNA methyltransferase. Family of enzymes that transfer the methyl group from S-adenosyl methionine (SAM) to DNA. In mammals there are three DNA methyltransferases: DNMT1, DNMT3a and DNMT3b.
DNMT3a and DNMT3b are de novo methyltransferases, they recognize something in the DNA that allows them to newly methylate cytosines. These are expressed mainly in early embryo development and they set up the pattern of methylation. DNMT1 is a maintenance methyltransferase and adds methylation to DNA when one strand is already methylated. These work throughout the life of the organism to maintain the methylation pattern that had been established by the de novo methyltransferases.
DNase I hypersensitive sites (DHS). are generic markers of regulatory DNA and contain genetic variations associated with diseases and phenotypic traits. A universal feature of active cis-regulatory elements—promoters, enhancers, silencers, chromatin insulators or enhancer blockers, and locus control regions—is focal alteration in chromatin structure triggered by binding of transcription factors (TFs), which supplants a canonical nucleosome and renders the underlying DNA accessible to nucleases and other protein factors. For more than 40 years, DHSs have provided reliable signposts for high-precision delineation of regulatory DNA in complex genomes. DHSs typically mark compact (less than 250 base pair (bp)) elements, and their appearance over a cis-regulatory region signifies its actuation (readying for activation), which may occur before, or coincident with, its functional activation. DHS mapping thus provides a generic tool for illuminating both active and potential regulatory landscapes (Meuleman 2020).
DNA sequencing. Determining the nucleotide sequence (primary structure) of DNA.
Sanger method or dideoxy DNA sequencing. Used from about mid 1970s. Four parallel solutions of a single-stranded DNA strand are mixed with DNA polymerase, the four dNTPs, and each solution with a low concentration of one of four types 2', 3'- dideoxynucleotides. When a dideoxynucleotide is build into the chain, the DNA synthesis is stopped, because the lack of a hydroxyl group at the 3' position of the pentose. Size-fractionating the four solutions on a polyacrylamide gel result in four columns from which the reverse complement nucleotide sequence of the single-stranded DNA can be read.
Automated Sanger or dideoxy DNA sequencing. In this case the four types of dideoxynucleotides are labeled with four different fluorescent dyes. Using only one solution with all reagents, the solution is put on a gel and submitted to electrophoresis. The moving bands pass a laser and a detector records the color of the band, which event is stored electronically.
Next-generation sequencing (NGS). (Metzker 2009, Goodwin 2016) From the early 2000s several technologies were developed to reduce cost and increase speed of the sequencing process. The following table summarises some features of several, mostly commercial, sequencing platforms.
| Template preparation | NGS chemistry | Platform | current company | |
| Emulsion PCR |
Single nucleotide addition (SNA) pyrosequencing |
Roche/454 | ||
| cleavable probe SBL |
SOLiD (Sequencing by Oligonucleotide Ligation Detection) |
ThermoFisher | ||
| non-cleavable probe SBL | Polonator | open source | ||
| Solid phase amplification | 3'- blocked reversible terminator | Illumina/ Solexa | Illumina | |
|
Single- molecule |
primer or template immobilization |
3'- unblocked reversible terminator | Helicos BioSciences | SeqLL |
|
polymerase immobilization |
real-time | Pacific Biosciences | Pacific Biosciences | |
Domain. See protein domain
Encyclopedia of DNA Elements (ENCODE) Consortium. "is an ongoing international collaboration of research groups funded by the National Human Genome Research Institute (NHGRI). The goal of ENCODE is to build a comprehensive parts list of functional elements in the human genome, including elements that act at the protein and RNA levels, and regulatory elements that control cells and circumstances in which a gene is active."
Short history:
2003 introduction ENCODE and pilot phase, 44 regions—approximately 1% of the human genome— where searched for functional elements in a few human cell lines
2007 publication of Pilot Phase results (ENCODE Project Consortium et al 2007) and start production scale-effort of the ENCODE project.
2012 publication of Second Phase (ENCODE 2) (ENCODE Project Consortium et al 2012)
2020 publication of Third Phase (ENCODE 3) (ENCODE Project Consortium et al 2020)
Enhancer. Region that activates transcription, often in a temporally and spatially restricted manner, by acting on a promoter. Enhancers can be located far from target promoters and are orientation independent.
Super-enhancers are large clusters of transcriptional enhancers that drive expression of genes that define cell identity. Disease-associated variation is especially enriched in the super-enhancers of disease-relevant cell types. Cancer cells generate super-enhancers at oncogenes and other genes important in tumor pathogenesis. Thus, super-enhancers play key roles in human cell identity in health and in disease (Hnisz 2013).
see ENCODE reference epigenome
see International Human Genome Consortium (IHEC)
see Roadmap Epigenomics Project
European Molecular Biology Laboratory (EMBL). centre of excellence for Europe's leading molecular biologists, provides life science researchers in Europe and beyond with access to the very latest in scientific technologies, infrastructure, and data resources.
Ensembl genome browser for vertebrate genomes that supports research in comparative genomics, evolution, sequence variation and transcriptional regulation. Ensembl annotate genes, computes multiple alignments, predicts regulatory function and collects disease data. Ensembl tools include BLAST, BLAT, BioMart and the Variant Effect Predictor (VEP) for all supported species.
F - J
Fermentation. Any metabolic process that releases energy from a sugar or other organic molecule, does not require oxygen, an electrochemical gradient or an electron transport system (like cellular respiration), but uses only a substrate-level phosphorylation to produce ATP and uses an organic molecule as the final electron acceptor. After fermentation there is no net change in the oxidation state of carbon. For instance in the conversion of glucose (C6H12O6) to lactate (C3H6O3), the H:C ratio remains the same.
Finngen is a large public-private partnership aiming to collect and analyse genome and health data from 500,000 Finnish biobank participants. FinnGen aims on one hand to provide novel medically and therapeutically relevant insights but also construct a world-class resource that can be applied for future studies. Major articles were published in Nature, volume 613, 19 januari 2023.
Folate/ folic acid (vitamin B9) 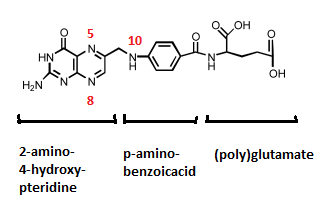
Definition
Many forms of folic acid, which itself is a synthetic molecule used as dietary supplement. Folates consists of three distinct functional groups:
1. a heterocyclic pteridine ring (2-amino-4-hydroxy-pteridine) that can be reduced or oxidized, and may bind one-carbon units
2. a para-amonobenzoicacid, that also may bind one-carbon units
3. a variable length polyglutamate tail
Folic acid can be reduced at nitrogen-8 by dihydrofolate reductase (DHFR) to dihydrofolate (DHF), which can be further reduced at nitrogen-5 by DHFR to tetrahydrofolate (THF). The enzyme serine hydroxymethyltransferase (SHMT) (pyridoxine dependent enzyme) can convert serine and THF to glycine and 5,10-Methylenetetrahydrofolate (5,10-CH2-THF), which can be part of several biochemical reactions, e.g. conversion to:
a. 5-Methyltetrahydrofolate (5-mTHF) by methylenetetrahydrofolatereductase (MTHFR) (Riboflavin dependent).
b. 5,10-Methenyltetrahydrofolate (5,10-CH=THF) by methylenetetrahydrofolate dehydrogenase 2 (MTHFD2). 5, 10-CH=THF can be further metabolised to 10-formyl-tetrahydrofolate (10-f-THF), which plays a role in purine metabolism.
Biological functions
Fragment template A fragment library is prepared by randomly shearing genomic DNA into small sizes of <1 kb and requires less DNA than would be needed for a mate-pair template (Metzker 2009).
G protein–coupled receptor (GPCR) large group of cell surface receptors, also called seven-transmembrane (7TM) receptors, because each consists of a single proteine with an extracellular N-terminus, an intracellular C-terminus and seven hydrophobic transmembrane domains (TM1-TM7) linked by three extracellular loops (ECL1-ECL3) and three intracellular loops (ICL1-ICL3). The transmembrane proteine is coupled to a heterotrimeric guanine nucleotide-binding proteine (G proteine). The G proteine consists of an α (Gα), β (Gβ) and γ (Gγ) subunit.
| Gq | Gi | Gs | |
|
signal transduction |
Activation phospholipase C (PLC) | Inhibition of adenylate cyclase | Activation of adenylate cyclase |
|
(receptor examples) Adrenergic
Muscarinic
Histamine
Serotonergic
GABA
Dopamine |
α1
M1 M3 M5
H1
5-HT2
|
α2
M2 M4
H3 H4
5-HT1 5-HT5
GABAB
D2 D3 D4 |
β1 β2 β3
H2
5-HT4 5-HT6-7
D1 D5 |
Gene Ontology (GO). resource that provides a computational representation of our current scientific knowledge about the functions of genes (or, more properly, the protein and non-coding RNA molecules produced by genes) from many different organisms, from humans to bacteria. It is widely used to support scientific research, and has been cited in tens of thousands of publications.
Genome aggregation database (gnomAD). "is a coalition of investigators seeking to aggregate and harmonize exome and genome sequencing data from a variety of large-scale sequencing projects, and to make summary data available for the wider scientific community." Major articles were published in Nature, vol 581, 28 may 2020.
see Ensembl genome browser (vertebrate)
see Ensembl Genomes (non-vertebrate)
Gluconeogenesis. Metabolic pathway for synthesis of glucose from non-carbohydrate precursors. In humans gluconeogenesis primarily occurs in liver, secundary in kidney. The main function is to provide cells who are dependent on glucose for energy production (brain, erythrocytes) with glucose during fasting and starvation.
This pathway shares the 7 reversible enzymes of glycolysis, but for the 3 irreversible enzymes gluconeogenesis has to have its own enzymes:
1. Pyruvate -> phosphoenolpyruvate (PEP). This step requires two reactions, and can occur in two ways:
A. Pyruvate is transported into mitochondrium, where the enzyme pyruvate carboxylase (coenzyme biotin or vitamine B7) converts it to oxaloacetate (using one ATP). Oxaloacetate has to be passed through the Malate-Aspartate Shuttle to the cytoplasm (because the mitochondrial membrane has no transporter for oxaloacetate). The cytosolic oxaloacetate is then converted by PEP carboxykinase (using one GTP) to PEP.
B. When lactate is the main source for pyruvate (Cori cycle), lactate is converted to pyruvate and NADH by lactate dehydrogenase. Pyruvate is again transported into mitochondrium, converted to oxaloacetate by pyruvate carboxylase, but this time oxaloacetate is immediately converted to PEP by mitochondial PEP carboxykinase to PEP, which can be transported to cytosol right away.
2. Fructose 1,6-biphosphate -> Fructose 6-phosphate by fructose-1,6-biphosphatase-1 (which forms a bifunctional enzyme with phosphofructokinase-1)
3. Glucose 6-phosphate -> Glucose by glucose 6-phosphatase. This enzyme is found on the lumenal side of the endoplasmic reticulum of hepatocytes, renal cells, and epithelial cells of the small intestine, so only these cells are able to supply glucose to the blood.
The net equation of gluconeogenesis is: 2 Pyruvate + 4ATP + 2GTP + 2NADH + 2H+ + 4H2O -> glucose + 4ADP + 2GDP + 6Pi + 2NAD+. Comparing this to the net equation of glycolysis makes the synthesis of glucose from pyruvate a relatively expensive process.
Glucose-6-phosphate dehydrogenase deficiency (G6PD deficiency). X-linked recessive disorder that results in defective glucoce-6-phosphate dehydrogenase enzyme, the first enzyme of the pentose phosphate pathway. It is also called "favism", but not all individuals with G6PD deficiency have favism, although all individuals with favism have G6PD deficiency. It is most frequent in populations in certain parts of Africa, Asia, the Mediterranean, and the Middle East. Prevalence is 7,5% in world population ranging to 35% in certain African areas, making it the most prevalent genetic disease ever known. The population distribution is related to heterozygote protective effect against Plasmodium falciparum and Plasmodium vivax malaria.
Glucose transporter. Membrane protein that facilitates transport of glucose across the plasma membrane.
Two main types are:
1. Sodium-glucose linked transporters (SGLTs). 14 types. Depend on the sodium concentration gradient. Present mainly on luminal surface intestinal cells and in renal tubules (glucose reabsorption).
2. Facilitated diffusion glucose transporters (GLUTs). 14 types divided into three classes. Class I comprises the best known GLUTs:
| Type |
Mnemonic |
Distribution | Notes |
| GLUT1 | BBB |
Blood (Erythrocytes) Baby (Fetus) Brain (endothelium Blood Brain Barrier) |
|
| GLUT2 | Kids Lips |
Kidney Liver Pancreas |
Bidirectional. In liver cells this allows uptake of glucose during glycolysis and release of glucose during gluconeogenesis. In pancreatic beta cells free flow of glucose (together with GLUT1) permits glucokinase to act as a glucose sensor, thus initiating insulin secreation when blood glucose is increasing. |
| GLUT3 | (are) Pink |
Placenta Neuron Kidney |
|
| GLUT4 |
Mother Father |
Muscle (Sketelal & Heart) Fat (Adipose tissue) |
Insulin dependent |
Glutamine (Gln, Q) proteinogenic amino acid.
The primairy functions of glutamine are to store nitrogen in muscle and to traffic it between organs (DeBernardinis 2010). Although it contributes only 4% of the amino acid in muscle protein, glutamine is the most abundant amino acid in plasma (> 20% of free amini acid) and more than 40% in muscle (90% of the body's glutamine store is in muscle).
Metabolic fates of glutamine (DeBernardinis 2010)
1. Reactions that use the γ-nitrogen
- Nucleotide biosynthesis. Glutamine is a required nitrogen donor for the de novo synthesis of purines and pyramidines.
- Hexosamine biosynthesis. Glutamine's amide group is transfered by glutamine:fructose-6-phosphate to fructose-6-phosphate to form glucosamine-6-phosphate, a precursor for N-linked and O-linked glycolsylation reactions.
2. Reactions that use the α-nitrogen or the carbon skeleton
These reactions use glutamate as substrate, so glutamine first has to be converted to glutamate by glutamase (GLS)
- Non essential amino acids
- Glutathione (GSH)
- Respiratory substrate. Oxidation of glutamine's carbon backbone in the mitochondria is a major fate of glutamine and a primary source of energy for proliferating cells, including lymfocytes, enterocytes, fibroblasts and some cancer cell lines. After conversion to glutamate, glutamate is converted to alpha-ketoglutarate and enters the citric acid cycle (anaplerosis). The carbon backbone leaves the citric acid cycle as malate (cataplerosis), is converted to oxaloacetic acid, PEP, pyruvate and finally reenters the citric acid cycle as acetyl-CoA.
- Ammoniagenesis
Gluten is the main storage protein of wheat grains. It is a complex mixture of hundreds of related but distinct proteins, mainly gliadin and glutenin, also refered to as prolamins, because they are characterized by high levels of proline and glutamine. Similar storage proteins are found in other cereals as zein in maize, secalin in rye, hordelin in barley, and avenin in oats. Collectively they are refered to as "gluten" (Biesiekierski 2017).
Gluten is classically defined as the largely proteinaceous mass which remains when dough made from wheat flour and water is gently washed in an excess of water or dilute salt solution to remove most of the starch and soluble material. The remaining "rubbery" material comprises about 75-80% protein on a dry mater basis (Shewry 2019).
A broad classification of seed proteins was described by (Osborne 1909) by using a sequential extraction process (called "Osborne fractionation") which distinghuished four fractions (Shewry 2019):
| Solvent | Fraction name | Notes |
| Water | albumins | |
| Dilute saline | globulins | |
| 60-70% alcohol | prolamins2 | Electrophoresis separates 4 groups: α-gliadins, β-gliadins, γ-gliadins and ω-gliadins |
| other1 | glutenins | The major glutenin components are in fact high molecular mass prolamin subunits which are stabilized by inter-chain disulphide bonds. By SDS-PAGE these can be separated into HMW (high molecular weight) and LMW (low molecular weight) subunits. LMW subunits can be further sub-divided into a major group B-type LMW subunits and two minor groups C-type and D-type LWM subunits. |
1 insolable or may be extracted in alkali
2 specific names in different cereals, in wheat gliadins etc
Comparisons of amino acid sequences of prolamin proteins distinguises the following groups (Shewry 2019):
1. HMW prolamins (HMW glutenins)
2. S-rich prolamins (α-gliadins, γ-gliadins, LMW glutenins)
3. S-poor prolamins (ω-gliadins)
A graphical representation of the relations between several prolamin subgroups was shown by (Bromilow 2017):
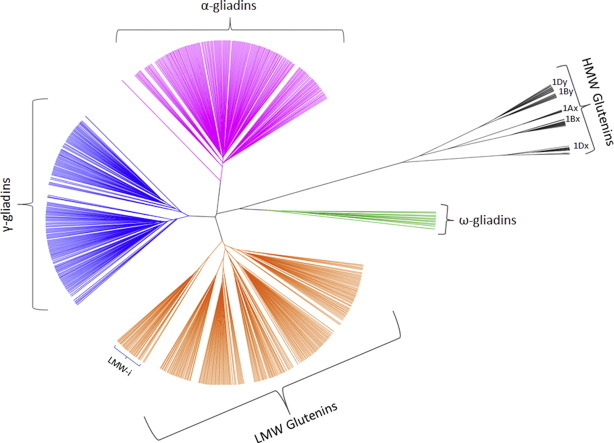
Gluten-Related Disorders (GRD)
Glycolysis. Anaerobic pathway located in the cytosol, that converts glucose into pyruvate and a hydrogen ion. In the first part of the pathway two ATP molecules are invested. Then glucose is split into two 3C molecules. In the last fase two ATP are gained per 3C molecule (total four ATP). So net gain per molecule glucose is two molecules ATP.
The net equation of glycolysis is: Glucose + 2 NAD+ + 2 ADP + 2 Pi -> 2 pyruvate + 2 NADH + 2 H+ + 2 ATP + 2 H2O
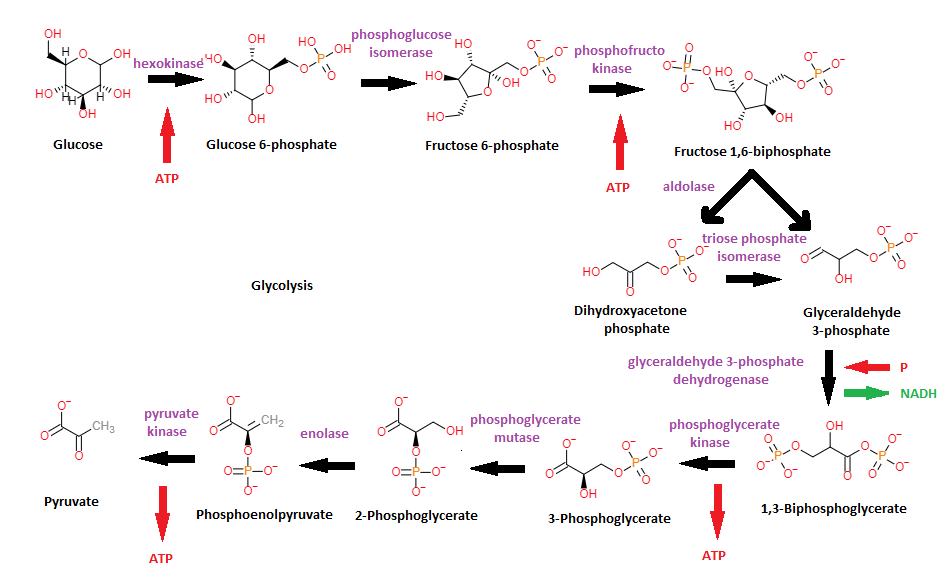
Seven of the ten reactions of glycolysis are reversible. These seven reactions, going in the other direction, and their enzymes, are also part of gluconeogenesis. The other three reactions are irreversible, and have to be bypassed with other enzymes in gluconeogenesis. The enzymes of these three reactions are important as sites of regulation of glycolysis. The three reactions and their enzymes are:
1. The first step of glycolysis: Glucose -> Glucose 6-phosphate catalyzed by hexokinase. The negative charge of phosphate prevents glucose from leaving the cell. Glucose 6-phosphate can also enter other pathways, like the pentose phosphate pathway and glycogen synthesis.
In the human genome four isoenzymes of hexokinase are encoded, hexokinase I to IV:
- hexokinases I, II and III have a high affinity for glucose (they have a low Km for glucose) and are inhibitited by their product glucose 6-phosphate.
- hexokinase IV, also called glucokinase, has a Km 100 times greater than the other hexokinases (it acts in the physiological range of blood glucose concentration) and it is not inhibited by its product glucose 6-phosphate. Hexokinase IV is present in liver, pancreas, small intestine and hypoyhalamus. It plays an important regulatory role in carbohydrate metabolism (Lenzen 2014). Glucokinase regulation is different in hepatocytes and pancreatic beta cells:
| Glucokinase (GK) | Hepatocyte | Pancreatic beta cell |
| Function | postprandial phosphorylation glucose for synthesis and storage glycogen | Sensor for glucose-induced insulin secretion |
|
> 48 hr starvation reduction activity reduction gene expression |
67% 100% |
50% 50% |
| refeeding restoration activity & gene expression | Insulin dependent | Glucose dependent |
| Inhibition | By binding to Glucokinase Regulatory Protein (GRP) GK is trapped in the nucleus. Fru-6-P strengthens this binding. Glucose and Fru-1-P weakens binding of GK to GRP, releasing it to the cytosol. | There is no expression of GRP in beta cells. GK is inhibited by ubiquitination and proteins like Midnolin and Parkin with ubiquitin-like domains. These interactions are weaker than the action of GRP |
| Activation by FBPase-2 | Binding of GK to the FBPase-2 part of the bifunctional PFK-2/FBPase-2 enzyme increases capacity of phosphorylating glucose without changing Km for glucose | |
2. The third step of glycolysis: Fructose 6-phosphate -> Fructose 1,6-biphosphate catalyzed by phosphofructokinase-1 (PFK-1). This is the commited step of glycolysis. Phosphofructokinase-1 forms a bifunctional enzyme with fructose-1,6-biphosphatase-1 (which converts Fructose 1,6-biphosphate to Fructose 6-phosphate, and is part of gluconeogenesis): PFK-1/FBPase-1. PFK-1 is inhibited allosterically by ATP and citrate, and stimulated by ADP. A small part of Fructose 6-phosphate is converted by the bifunctional enzyme PFK-2/FBPase-2 into fructose 2,6-biphosphate. Fru 2,6-BP stimulates PFK-1 and inhibits FBPase-1, playing an important part in switching between glycolysis and gluconeogenesis. Glucagon stimulates cAMP mediated phosphorylation of PFK-2/FBPase-2, activating FBPase-2, so gluconeogenesis is stimulated. Insulin stimulates dephosphorylation of PFK-2/FBPase-2, activating PFK-2, so stimulating glycolysis.
The second function of PFK-2/FBPase-2 is that the FBPase-2 domain is a strong activator of glucokinase.
3. The last step of glycolysis: Phosphoenolpyruvate (PEP) -> Pyruvate catalysed by pyruvate kinase.
Histone. Protein involved in packaging of DNA. They have a very important role in the control of chromatin structure and thus in the regulation of gene transcription. Histones are rich in arginine and lysine, amino acids with positive charged side chains. The two kinds of histones are:
1. Core histones: H2A, H2B, H3 and H4
2. Linker histones H1/H5
In the core histones two elements can be recognized:
1. The histone fold
2. The N terminal tail
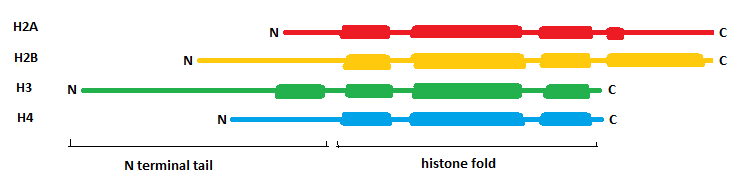
1. The histone fold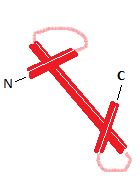
This is a structural motif shared by the four core histones: three alpha helices connected by two loops. They fold as can be seen on the right. Histones 2A and 2B form a dimer by an interaction called the "handshake". The same happens with histones H3 and H4. Two dimers of each form a octamer, the nucleosome, around which DNA is wound.
2. N terminal tail
Homology, sequence. is the biological homology between DNA, RNA or protein sequences, defined in terms of shared ancestry in the evolutionary history of life.
Terms and definitions (Koonin 2005):
1. Homologs: Genes sharing a common origin
Orthologs: Genes originating from a single ancestral gene in the last common ancestor of the compared genomes.
Pseudoorthologs: Genes that actually are paralogs but appear to be orthologous due to differential, lineage-specific gene loss.
Xenologs Homologous genes acquired via xenologous gene displacement (XGD) by one or both of the compared species but appearing to be orthologous in pairwise genome comparisons.
Co-orthologs Two or more genes in one lineage that are, collectively, orthologous to one or more genes in another lineage due to a lineage-specific duplication(s).
Members of a co-orthologous gene set are inparalogs relative to the respective speciation event.
2. Paralogs: Genes related by duplication
Inparalogs (symparalogs) Paralogous genes resulting from a lineage-specific duplication(s) subsequent to a given speciation event (defined only relative to a speciation event, no absolute meaning).
Outparalogs (alloparalogs) Paralogous genes resulting from a duplication(s) preceding a given speciation event (defined only relative to a speciation event, no absolute meaning).
Pseudoparalogs Homologous genes that come out as paralogs in a single-genome analysis but actually ended up in the given genome as a result of a combination of vertical inheritance and horizontal gene transfer (HGT).
Housekeeping gene. Gene that is required to maintain basic cellular function and so is typically expressed in all cell types of an organism. Some housekeeping genes are transcribed at a relatively constant rate and these genes can be used as a reference point in experiments to measure the expression rates of other genes.
Human accelerated regions (HARs). ancestrally conserved genomic regions (regions that are highly evolutionarily conserved among many mammals) that show a significantly accelerated rate of substitution in the human lineage since divergence from our common ancestor with the chimpanzee (Pollard 2006). Pollard et al reported 49 of such regions (HAR1-HAR49) in 2006. All but two HARs lie outside protein-coding sequences, but they often lie near protein-coding genes that have neurodevelopmental functions. The most rapid change was found in HAR1, which lies in the last band of chromosome 20q and is part of a pair of overlapping divergently transcribed non-coding RNA genes, HAR1F (Human Accelerated Region 1, Forward, also called HAR1A) and HAR1R (Human Accelerated Region 1, Reverse, also called HAR1B). HAR1F was found to be expressed at 7 to 19 gestational weeks in Cajal-Retzius cells in the dorsal telencephalon. HAR1F is also expressed in the ovary and testis of adult humans.
Human Cell Atlas. project aimed to map all the cells in the human body so as to facilitate a better understanding of human heath, as well as research in diagnosis, monitoring and treatment of disease. White Paper
Human genes and genetic diseases, databases
see GeneCards
see Genetic Association Database (GAD) frozen as of 09/01/2014 but still downloadable
see Online Mendelian Inheritance in Man (OMIM)
Human genome. Total of human DNA nucleotide sequences.
Composition of the human genome:
| Percentage | Component | Description |
| 26 | Introns | Intragenic region. Any nucleotide sequence within a gene that is removed by RNA splicing |
| 20 | LINEs | Long Interspersed Nuclear Element. Transposable element |
| 13 | SINEs | Short Interspersed Nuclear Element. Transposable element |
| 8 | LTR retrotransposons | Long Terminal Repeats retrotransposons. Transposable element |
| 5 | segmental duplications | long DNA sequences that have nearly identical sequences (90-100%) and exist in multiple locations as a result of duplication events. |
| 3 | DNA transposons | Transposable element |
| 3 | simple sequence repeats | a DNA tract consisting of a relatively short base-pair motif that is repeated several to many times in tandem |
| 1,5 | protein coding genes | |
| 20 | other | miscellaneous heterochromatin and unique sequences |
Inflammation complex set of interactions among soluble factors and cells that can arise in any tissue in response to traumatic, infectious, post-ischemic, toxic or autoimmune injury. The "inflammation process" includes a tissue-based startle reaction to trauma; go/no-go decisions based on integration of molecular clues for tissue penetration by microbes; the beckoning, instruction and dispatch of cells; the killing of microbes and host cells they infect; liquefaction of surrounding tissue to prevent microbial metastasis; and the healing of tissues damaged by trauma or by the host's response (Nathan 2002).
Inflammation can be seen as a system of information flow in response to injury and infection. The host reacts to trauma as if there is infection, until proven otherwise. If there is infection, the goal is to prevent its spreading, even at the cost of further tissue damage. If the spread of infection is checked, the inflammation reaction has to be decelerated, and repair of damaged tissue started (Nathan 2002).
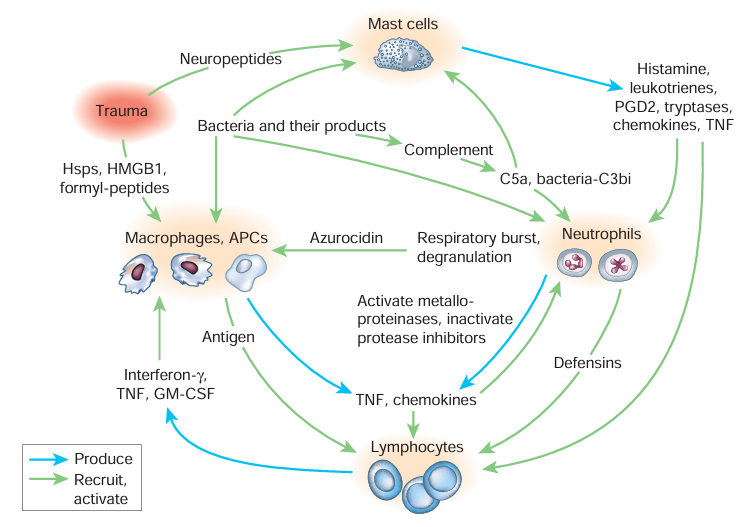
Information flow in the early stages following mild trauma with infection. Each cell commits to recruit and activate others based on multiple inputs, generally requiring evidence of both injury and infection, before it joins fully in amplifying the inflammatory process. Not shown are interactions among leukocytes, endothelium, platelets and coagulation factors; the generation of stop signals; and the flow of information over subsequent days, including the transition to wound healing (Nathan 2002).
Go signals in early checkpoints. Tissue damage causes three types of go signals:
1. bioactive peptides released by neurons in response to pain.
2. intracellular proteins released by broken cells. E.g. heat-shock proteins (HSP), transcription factor HMGB1 (high mobility group 1) and mitochondrial peptides bearing the N-formyl group characteristic of prokaryotic proteins.
3. products shed or secreted by microbes.
Mast cells can be considered "first responders" to these signals. The best characterized trigger for mast cells are allergens complexed to IgE. Fc receptors of mast cells are constantly coated with IgE. Crosslinking these receptors triggers degranulation. Microbes can trigger mast cells through Toll-like receptors (TLRs). Mast cells can also be activated by cytokines and chemokines, in particular tumour-necrosis factor-α (TNF-α) and monocyte chemoattractant protein 1 (MCP-1), which they also release themselves, so making way to positive feedback. Mast cells can release a very broad array of mediators and cell-cell signaling molecules:
1. fast release by exocytosis from plentyful intracellular stores (histamine, specific proteases from members of tryptase and chymase families, and TNF-α)
2. subsequent rapid synthesis of arachidonic acid, prostaglandines and leukotrienes (Benoist and Mathis 2002).
Integrated Microbial Genomes and Microbiomes system to support the annotation, analysis and distribution of microbial genome and microbiome datasets sequenced at DOE's Joint Genome Institute (JGI).
International Society for Phylogenetic Nomenclature (ISPN) was established to encourage and facilitate the development and use of, and communication about, phylogenetic nomenclature. Toward this end, the ISPN organizes periodic scientific meetings and oversees the implementation of a code of rules to govern phylogenetic nomenclature—the PhyloCode. Its current version is specifically designed to regulate the naming of clades.
International Union of Biochemistry and Molecular Biology (IUBMB) is devoted to promoting research and education in biochemistry and molecular biology throughout the world. Recommendations on Biochemical & Organic Nomenclature, Symbols & Terminology etc.
International Union of Pure and Applied Chemistry (IUPAC) is the world authority on chemical nomenclature and terminology, including the naming of new elements in the periodic table; on standardized methods for measurement; and on atomic weights, and many other critically-evaluated data. Recommendations on Organic & Biochemical Nomenclature, Symbols & Terminology etc.
International Wheat Genome Sequencing Consortium (IWGSC) is an international, collaborative consortium, established in 2005 by a group of wheat growers, plant scientists, and public and private breeders. The vision of the IWGSC is a high quality genome sequence of bread wheat that serves as a foundation for the accelerated development of improved varieties and that empowers all aspects of basic and applied wheat science.
K - O
Kyoto Encyclopedia of Genes and Genomes (KEGG). database resource for understanding high-level functions and utilities of the biological system, such as the cell, the organism and the ecosystem, from molecular-level information, especially large-scale molecular datasets generated by genome sequencing and other high-throughput experimental technologies.
Krebs cycle see Citric acid cycle
Malate-Aspartate Shuttle biochemical mechanism to transport reducing equivalents from cytosolic NADH across the semipermeable inner membrane, but impermeable for NADH, of the mitochondrion. 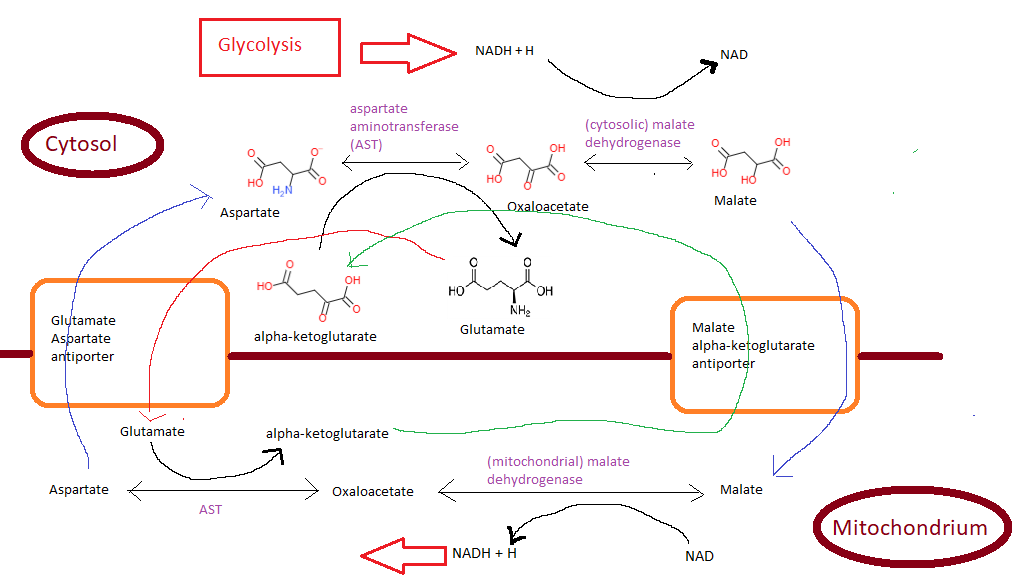
Mate-pair template A genomic library is prepared by circularizing sheared DNA that has been selected for a given size, such as 2 kb, therefore bringing the ends that were previously distant from one another into close proximity. Cutting these circles into linear DNA fragments creates mate-pair templates (Metzker 2009).
MEROPS the peptidase database. Information resource for peptidases (also termed proteases, proteinases and proteolytic enzymes) and the proteins that inhibit them. The Summary page describing a given peptidase can be reached by use of an index under its Name, MEROPS Identifier or source Organism. The Summary describes the classification and nomenclature of the peptidase and offers links to supplementary pages showing sequence identifiers, the structure if known, literature references and more. The MEROPS database uses an hierarchical, structure-based classification of the peptidases. In this, each peptidase is assigned to a Family on the basis of statistically significant similarities in amino acid sequence, and families that are thought to be homologous are grouped together in a Clan. There is a Summary page for each family and clan, and these again have indexes. Each of the Summary pages offers links to supplementary pages.
Methionine (met, M) proteinogenic amino acid.
Metzincin large gene superfamily of zinc protease.
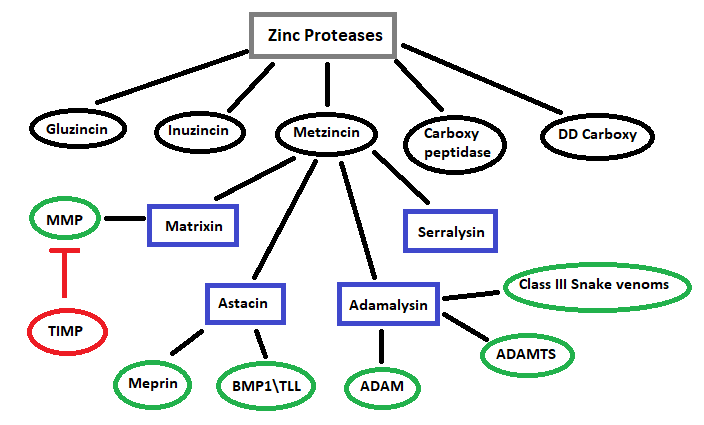 Schematic representation of subdivisions within the Metzincin gene superfamily, MMP, Matrix metalloproteinases. TIMP, Tissue Inhibitor of Metalloproteinases. ADAM, A Disintegrin And Metalloproteinase. ADAMTS, A Disintegrin And Metalloproteinase with Thrombospondin motifs. BMP1/TLL, Bone Morphogenetic Protein/ Tolloid Like (Huxley-Jones et al 2007).
Schematic representation of subdivisions within the Metzincin gene superfamily, MMP, Matrix metalloproteinases. TIMP, Tissue Inhibitor of Metalloproteinases. ADAM, A Disintegrin And Metalloproteinase. ADAMTS, A Disintegrin And Metalloproteinase with Thrombospondin motifs. BMP1/TLL, Bone Morphogenetic Protein/ Tolloid Like (Huxley-Jones et al 2007).
Microbial Genome Database for Comparative Analysis database for comparative analysis of completely sequenced microbial genomes, the number of which is now growing rapidly. The aim of MBGD is to facilitate comparative genomics from various points of view such as ortholog identification, paralog clustering, motif analysis and gene order comparison. References: Nucleic Acids Res. 47:D382-D389 (2019)
Model organism. Species that has been chosen for biological study to optimize limited research resources, because it has some characteristics that makes it attractive to study certain biological phenomena, in combination with the fact that the species is easy to handle and convient to work with.
(Goldstein & King 2017) discuss that increasingly traditional model organisms are not the best choice to answer a research question. For instance some evolutionary cell biology questions specifically require the study of organisms at key places on the tree of life: "When thinking about cell biological mechanisms, or indeed any biological phenomena, the quest to indentify universal principles benefits from an understanding of evolutionary history. For example, membrane trafficking is solely a phenomenon of eukaryotic biology; investigating its origins provides a valuable complementary approach for identifying key regulatory mechanisms. The traditional model organisms are all members of a recently derived group sometimes referred to as the "crown eukaryotes" and provide only a narrow window into the evolution of membrane trafficking".
Some examples of model organisms
|
Species
|
Common name
|
Type
|
Genome size (kilobase) Number of genes
|
Model organism database
|
|
Escherichia coli
|
Bacteria
|
4639 4289
|
|
|
|
Saccharomyces cerevisiae
|
Baker's yeast
|
Yeast
|
12,069 ~6300 |
|
|
Arabidopsis thaliana
|
Thale cress Zandraket
|
Flowering plant
|
~142,000 ~26,000 |
|
|
Caenorhabditis elegans
|
Nematode
|
~97,000 ~20,000 |
|
|
|
Drosophila melanogaster
|
Fruitfly
|
Insect
|
~137,000 ~14,000 |
|
|
Danio rerio
|
Zebrafish |
Fresh water fish
|
1,412,464 26,206 |
|
|
Xenopus laevis
Xenopus tropicalis
|
African clawed frog
Western clawed frog
|
Frog
|
|
|
|
Mus musculus
|
House mouse
|
Mammal
|
2.8 x 106 ~23,000
|
Mouse Ageing Cell Atlas
|
|
Rattus norvegicus
|
Brown rat
|
Mammal
|
|
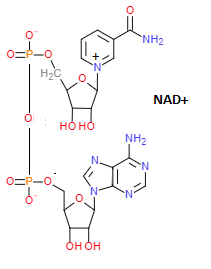
Nicotinamide adenine dinucleotide. Electron carrying cofactor central to metabolism, consisting of two nucleotides, one containing adenine, the other nicotinamide. It exists in two forms: oxidized (NAD+) and reduced (NADH).
Another important electron carrier is the closely related NADP+/NADPH. Cell metabolism thus has two independant electron carriers, NAD being used for catabolic processes producing ATP, and NADP for anabolic processes. The cell is maintaining a high NAD+/NADH ratio and a low NADP+/NADPH ratio.
Precursors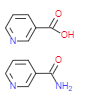
- Niacine = Nicotinic acid (NA)
vitamin B3 complex - Nicotinamide (NAM)
- Nicotinamide riboside (NR)
Tryptophan
Moiety (functionele groep) part of a molecule that is given a name because it is identified as a part of other molecules as well.
Nixtamalization. process for the preparation of maize (corn), or other grain, in which the corn is soaked and cooked in an alkaline solution, usually limewater (but sometimes woodash lye), washed, and then hulled. In industrial settings enzymatic nixtamalization with a protease is used. Nixatamalization has several benefits:
- maize can be more easily grounded and it improves flavor and aroma
- reduces commonly present carcinogenic mycotoxins by more than 90 %
- reduces amount of the protein zein, so total amount of protein is reduced, but balance among essential aminoacids is improved
- increases bio-availability of niacin, thus reducing the incidence of pellagra
Nuclease. Enzyme capable of cleaving the phosphodiester bonds between the nucleotides of nucleic acids.
There is no simple way to classify nucleases (Yang 2011):
- Nucleases can be proteins or RNA (ribozyes)
- They can be DNases, RNases or both
- Exonucleases cleave one nucleotide at a time from the end of a nucleic acid. They can be further divided in acting from 3' to 5' or 5' to 3'.
Endonucleases cleave the phosphodiester bond within a nucleic acid. Some endonucleases cleave DNA in a nonspecific way (regardless of nucleotide sequence). Many endonucleases, called restriction endonucleases, cleave only at very specific nucleotide sequences.
Nucleoside/Nucleotide A nucleoside is a molecule with a nitrogenous base and a five-carbon sugar. Nucleotides also have one or more phosphate groups.
Mnemonic: "S(ide has no phosphate) -> T(ide has phosphate)"
Nucleosides and nucleotides are named after the nitrogenous base. The bases are either a pyrimidine or a purine.
Mnemonic: "CUT (cytosine, uracil, thymine) the pyramids"
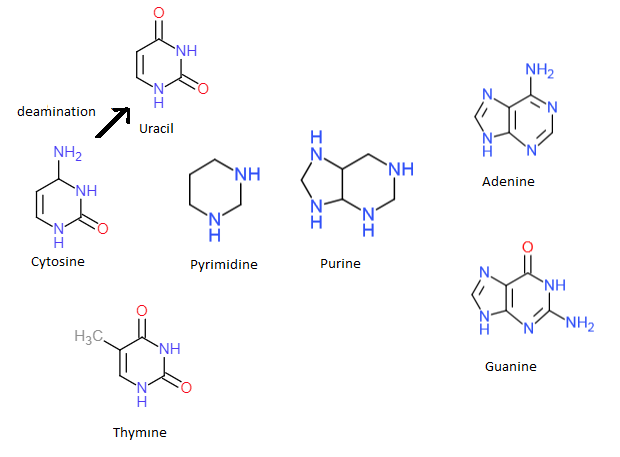
Nucleotides and nucleosides have a five-carbon sugar (pentose), which can be either ribose or deoxyribose:
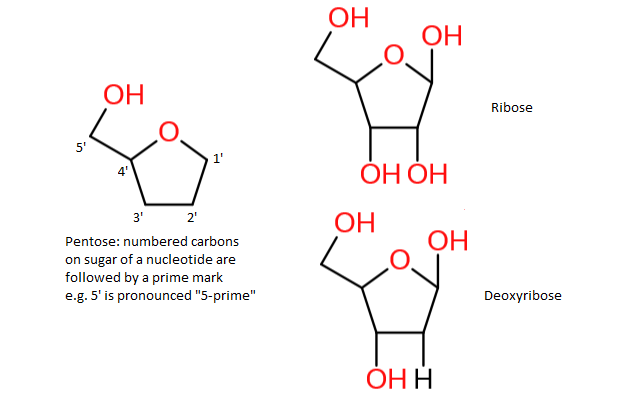
Nucleotides have one to three phosphate groups connected tot the 5' carbon of the pentose. Generally speaking we have NMP (nucleotide monophosphate), NDP (nucleotide diphosphate) and NTP (nucleotide triphosphate). For example: the nucleotide adenosine triphosphate (ATP):
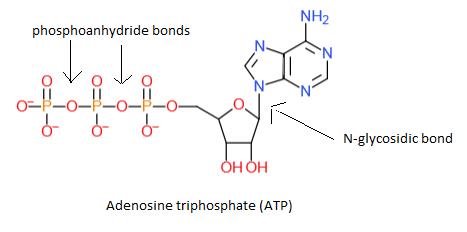
Biological functions of nucleotides: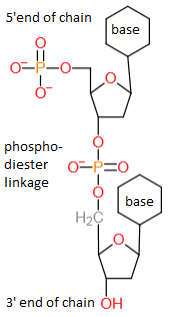
1. They are the subunits of nucleic acids, forming long molecules by phosphodiester links between the 5' and 3' carbon atoms. DNA is made from deoxyribose, RNA is made from ribose nucleotides. In DNA in the two strands paring exists between cytosine and guanine, and thymine and adenine. In RNA thymine is replaced by uracil (which is an unmethylated form of thymine). Notice that cytosine is very unstable and can change into uracil (spontaneous deamination). Furthermore cytosine can be methylated by methyltransferase into 5-methylcytosine.
Here we see two deoxyribosenucleotides with a phophodiester linkage:
In vivo nucleic acids can only be synthesized from the 5' to the 3' direction.
Note that in some laboratory procedures, like DNA sequencing, dideoxy nucleotides (the 3' hydroxyl group is replaced by a hydrogen atom, which when build into the nucleic acid, prevent further growth of the nucleic acid chain) and other modified nucleotides are used.
2. Adenosine triphosphate (ATP), see above, is the most important energy carrier in cells. Hydrolysis of a phosphoanhydride bond of ATP gives ADP, a phosphategroup and energy.
3. They are part of several activated carrier molecules that play an important role in metabolism. For instance:
Carriermolecule carried group
- ATP phosphate
- NADH, NADHP electrons and hydrogens
- Acetyl coA acetyl group
- S-Adenosylmethionine methyl group
- Uridine diphosphate glucose glucose
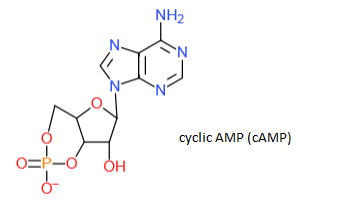
4. They can be used in cell signaling. For example:
Open Reading Frame (ORF). One of three possible reading frame's in a nucleotide sequence that begins with a start codon (ATG in DNA, AUG in RNA) and ends with a stop codon (TAG, TAA of TGA in DNA, UAG,UAA or UGA in RNA).
Open Reading Frame Finder: https://www.ncbi.nlm.nih.gov/orffinder/
Osborne, Thomas Burr (August 5, 1859 – January 29, 1929), biochemist. Independently discovered Vitamin A. Shewry(2002) calls him "the father of of plant protein chemistry". (Osborne 1909): "The vegetable proteins" (monograph).
Oxidative phosphorylation Last stage of cellular respiration taking place in mitochondria.
P - T
Pan-Cancer Analysis of Whole Genomes (PCAWG) study is an international collaboration to identify common patterns of mutation in more than 2,600 cancer whole genomes from the International Cancer Genome Consortium. Major articles were published in Nature, vol 578, 6 february 2020.
Pathguide. contains information about 702 biological pathway related resources and molecular interaction related resources.
PathVisio free open-source pathway analysis and drawing software which allows drawing, editing, and analyzing biological pathways. It is developed in Java and can be extended with plugins.
Pentose phosphate pathway (PPP) or hexose monophosphate shunt. Cytoplasmic pathway which metabolizes glucose-6-phosphate to some specialised products needed by cells:
- NADPH, which is needed for
* fatty acid synthesis (liver, adipose, lactating mammary gland)
* synthesis of cholesterol and steroid hormones (liver, adrenal glands, gonads)
* oxidative stress homeostasis (cells which are directly exposed to oxygen: erythrocytes and the cells of the lens and cornea)
- Ribose-5-phosphate which is necessary for the production of nucleotides and nucleic acids (rapidly dividing cells, such as those of bone marrow, skin, intestinal mucosa, and tumors)
- Products needed in the Shikimate pathway in bacteria, archaea, fungi, algae, some protozoans and plants for the biothsynthesis of folates and aromatic amino acids (tryptophan, phenylalanine, tyrosine)
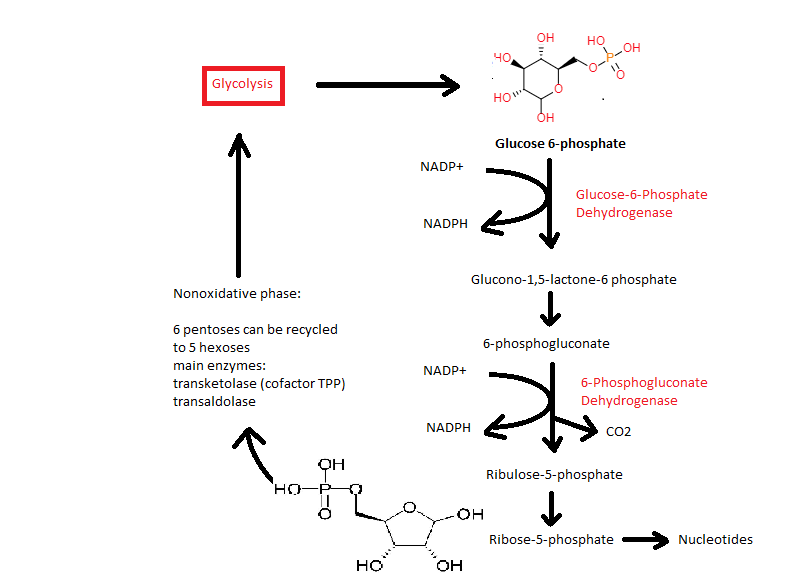
Peptidase also protease, proteinase or proteolytic enzyme. Enzyme that catalyses proteolysis, the breakdown of proteines into smaller polypeptides or proteinogenic amino acids.
Peto's paradox: between species, contrary to inside species, there seems to be no correlation between incidence of cancer and body mass (number of cells) or life expectancy (number of cell divisions). Carcinogenesis is supposed to correlate with accumulation of mutations during cell division, so the more cells (body mass) or the more cell divisions (age) the higher the incidence of cancer. Peto's paradox, if true, is yet not explained. It is often postulated that natural selection on large size or extended longevity is inherently inseparable from the evolution of anticancer defenses. Vincze et al (2022) found the most robust support for Peto's paradox till now, using data from large numbers of species and individuals from mammals living in zoos. They also found the lowest cancer mortality in Artiodactyla (even-toed ungulates, like pigs, hippopotamuses, antelopes, deer, giraffes, camels, llamas, alpacas, sheep, goats and cattle). The highest cancer mortality risk was found in Carnifora. Also vertebrate, but not invertebrate, diet correlated with cancer mortality. Mammals frequently consuming mammalian prey showed highest cancer mortality.
Several explanations were explored:
- use of hormonal contraception (which should show sex bias, which was not found)
- high-fat, low-fibre diet
- top of the food chain effect due to accumulation of carcinogenic compounds (does not explain differences in invertebrate and vertebrate diets)
- exposure to oncogenic pathogens (is consistent with highest cancer risk in mammals eating mammals)
- other: low microbiome diversity, limited exercise
Promoter. Region that is located immediately upstream of a protein-coding gene, and binds to RNA polymerase II; where transcription is initiated.
In the case of Pol II, one class of human promoters contains CpG islands that can impair the assembly of inhibitory nucleosomes and facilitate polymerase access. Promoters such as these are often found at housekeeping genes that encode for proteins that are required in all the cell types of an organism. The activity of promoters that contain CpG islands can be altered by DNA methylation.
Another class of Pol II promoters contains a TATA element upstream of the transcription start site; promoters of this class are often found at genes that are cell-type-specific and regulated during differentiation (Cramer 2019).
Protein databases and tools. (Xu 2004), (Koehl 2006)
Protein Sequence Databases
| PIR | Protein Information Resource |
| Swiss-Prot | Manually annotated an reviewed section of the UniProt Knowledgebase (UniProtKB) |
| UniProt | resource of protein sequence and functional information |
Protein Structure Databases
| PDB |
Protein Data Bank. Repository of protein structures |
| PDB at a Glance |
Hypertext-based table of contents of the April 1994 release of the Brookhaven Protein Databank (PDB) |
| PDBe |
European Bioinformatics Institute interface to the PDB, with integration to EBI resources |
| PDBsum |
Pictorial database of 3D structures in the Protein Data Bank |
| PISCES |
Protein sequence culling server: generates subsets of PDB based on users’ criteria |
Protein Family Databases, Sequence-Based
| COG |
Clusters of Orthologous Group (focus on evolution) |
| Pfam |
large collection of protein families, each represented by multiple sequence alignments and hidden Markov models (focus on function) |
| ProDom |
comprehensive set of protein domain families automatically generated from the UniProt Knowledge Database (focus on sequence domain) |
| SMART |
Simple Modular Architecture Research Tool |
Protein Family Databases, Structure-Based
| CATH |
classification of protein structures downloaded from the Protein Data Bank |
| ECOD |
Evolutionary Classification of Protein Domains |
| SCOPe |
Structural Classification of Proteins — extended |
Protein Family Databases, Function-Based
| BRENDA |
Comprehensive Enzyme Information System |
| ENZYME |
repository of information relative to the nomenclature of enzymes |
| GPCRdb |
data, diagrams and web tools for G protein-coupled receptors (GPCRs) |
| InterPro | InterPro |
| KEGG |
Kyoto Encyclopedia of Genes and Genomes |
| REBASE |
Restriction Enzyme Database |
| TransportDB |
relational database describing the predicted cytoplasmic membrane transport protein complement for organisms whose complete genome sequences are available |
Protein domain. It is a well accepted idea that proteins can be thought of consisting of one or more recognizable "units" or "evolutionary building blocks" that may appear in a variety of different proteins. However it proves to be very difficult to give a consistent definition of domains. (Koehl 2006) mentions five working definitions:
(1) regions that display a significant level of sequence similarity;
(2) the minimal part of a protein that is capable of performing a function;
(3) a region of a protein with an experimentally assigned function;
(4) a region of a structure that recurs in different contexts in different proteins;
(5) a compact, spatially distinct unit of protein structure.
Differences between Motifs and Domains:
| Motif or Fold | Domain | |
| Definition | a recognizable folding pattern involving two or more elements of secondary structure and the connection(s) between them | a part of a polypeptide chain that is independently stable or could undergo movements as a single entity with respect to the entire protein |
| Structure | supersecondary (not part of the primary - quarternary hierachy) | tertiary |
| Stability | may or may not be stable | stable by itself |
| Formed by | the connected alpha-helices and beta-sheets through loops | the formation of disulfide bridges, ionic bonds and hydrogen bonds between amino acid side chains |
| Functional role | structural function mainly | functional unit of the protein |
| Functions through protein families | similar | unique |
| Examples |
Beta hairpin Helix-loop-helix Helix-turn-helix Zinc finger |
Basic Leucine Zipper Domain (bZIP domain) Immunoglobulin-like domains Zinc finger DNA-binding domain (ZnF_GATA) |
Proteine-Proteine Interaction (PPI). specific physical contacts between proteine pairs that occur by selective molecular docking in a particular biological context (De La Rivas 2010).
There are two groups of methods to get information about PPIs: computational and experimental. Each method has its limitations and is prone to produce false negative and false positive results. Results get more reliable by combining several methods. Often high-throughput methods are followed by using low-throughput methods.
1. Computational methods
| Method | Description |
| Phylogenetic profiling | If two or more proteins are concurrently present or absent across several genomes, then they are likely functionally related |
| Similar phylogenetic trees | Phylogenetic trees of receptors and ligands were observed to be more similar than due to random chance |
| Rosetta Stone or Domain Fusion method | Interacting proteines may be observed to have been fused into one single proteine in some genomes |
| Conserved gene neighborhood | If genes encoding two proteines are neighbors in many genomes, they may be functionally related |
PubMed comprises more than 33 million citations for biomedical literature from MEDLINE, life science journals, and online books. Citations may include links to full text content from PubMed Central and publisher web sites.
Pyruvic acid. Smallest of the alpha-keto acids. It is an intermediate in several metabolic pathways in the cell:
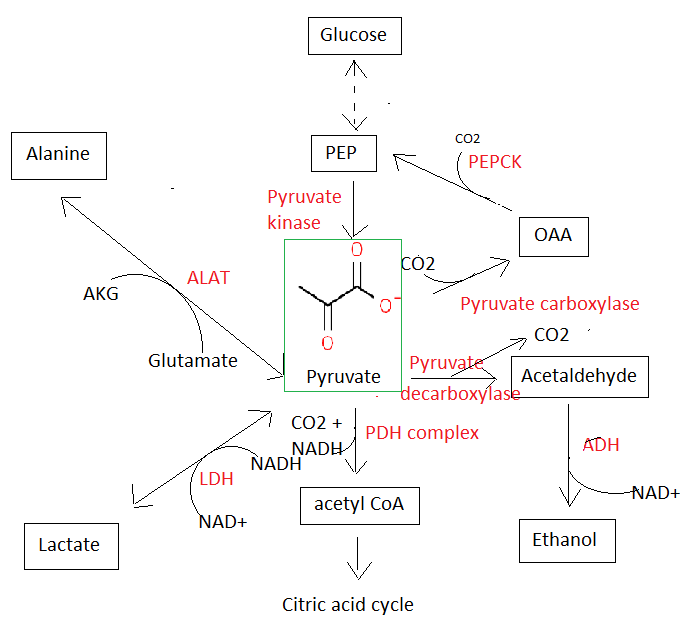
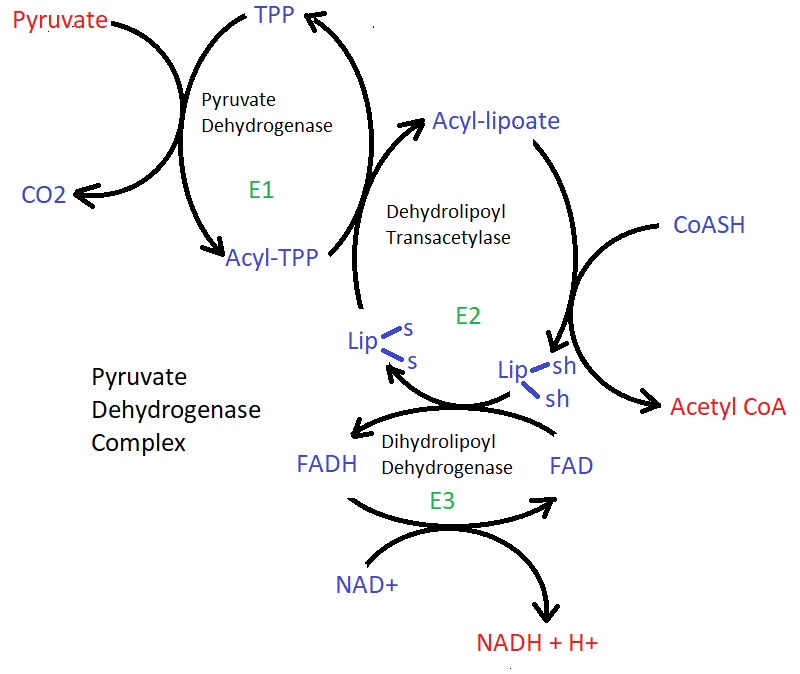
Pyruvate dehydrogenase complex (PDH complex). Complex of three enzymes and five coenzymes that convert pyruvate to acetyl CoA, and thus links glycolysis to the citric acid cycle.
| Enzyme | Cofactor | Comments | |
| E1 | Pyruvate dehydrogenase | thiamine pyrophosphate (TPP) | first decarboxylates pyruvate (forming CO2), then oxydizes the hydroxyethyl group to a acetyl group, the electrons reduce the disulphide of lipoate from E2, and the acetyl group is transferred to one of the -SH groups of the reduced lipoate. |
| E2 | Dehydrolipoyl transacetylase | lipoate, coenzyme A | the swinging lipoyllysyl arms form the centre of the PDH complex, the acetyl group from E1 is transfered to coenzyme A, forming acetyl coenzyme A, the electrons are passed to E3 |
| E3 | Dehydrolipoyl dehydrogenase | FAD, NAD | the electrons are first passed to FAD, regenerating the disulphide form of lipoate, then pass the electrons to NAD+, forming NADH and H+ |
Receptor (in biochemistry and pharmacology). Chemical structure, composed of protein, that binds the signal molecule (ligand) and initiates a process that amplifies the signal, integrates it with input from other receptors, and transmits the information throughout the cell (Nelson 2013).
Four major groups of receptors are:
|
Ligand-gated ion channels (ionotropic receptors) |
G protein-coupled receptors (GPCRs) (metabotropic receptors) |
Kinase-linked and related receptors |
Nuclear receptors | |
| Location | cell membrane | cell membrane | cell membrane |
cytoplasm (migrates to nucleus after ligand binding) |
| Effector | ionchannel | enzyme or ion channel | enzyme | gene |
| Timescale | milliseconds | seconds - minutes | minutes - hours | hours - days |
|
human drug targets by gene family1 |
19% | 12% | 10% | 3% |
|
% drugs targetting this group1 |
18% | 33% | 3% | 16% |
| Example receptors |
nicotinic acetylcholine GABAA 5-HT3 |
muscarinic acetylcholine GABAB 5-HT1-2 5-HT4-7 adrenaline |
insulin glucokinase |
steroid vitamin D |
| More detailed list | Ion channels | GPCRs | Kinases | Nuclear hormone receptors |
1Santos 2017. Drug target % > % drugs (Kinases) reflects few drugs effect lots of targets (polyfarmacology). The opposite means lots of drugs effect few targets (high selective pharmacoloy) (Nuclear receptors )
Reducing equivalent any of a number of chemical species which transfer the equivalent of one electron in redox reactions
Restriction. The cutting of DNA at specific sites.
Restriction endonuclease. Nuclease that cleaves the phosphodiester bond within a polynucleotide chain at or near specific recognition sites. Restriction endonucleases are important tools in molecular biology. REBASE http://rebase.neb.com/rebase/rebase.html is a database for restriction endonucleases.
RNA polymerase. Enzyme that synthesizes RNA from a DNA template. In eukaryotes there are several types of RNA polymerases:
Pol I produces the large ribosomal RNA precursor
Pol II synthesizes messenger RNAs and a variety of non-coding RNA
Pol III produces transfer RNAs and the small ribosomal RNA
In plant cells two additional Pol II-like polymerases are found which are involved in gene silencing:
Pol IV synthesizes siRNA
Pol V synthesizes RNAs involved in siRNA-directed heterochromatin
Scissile bond. (scissile = capable of being cut smoothly or split easily) covalent chemical bond that can be broken by an enzyme.
Sequencing is determining the primary structure of an unbranched biopolymer.
see DNA sequencing
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE)
Systems Biology Markup Language (SBML) a free and open interchange format, based on XML, for computer models of biological processes. SBML is useful for models of metabolism, cell signaling, and more.
Technology, molecular biological
Avidin-Biotin Interaction (ThermoFisher),
Avidin-Biotin Technical Handbook (ThermoFisher)
see DNA sequencing
Template This recombinant DNA molecule is made up of a known region, usually a vector or adaptor sequence to which a universal primer can bind, and the target sequence, which is typically an unknown portion to be sequenced. Current methods generally involve randomly breaking genomic DNA into smaller sizes from which either fragment templates or mate-pair templates are created (Metzker 2009).
Thiamine pyrophosphate (TPP or ThPP). Coenzyme formed from thiamine (vitamine B1). It plays an important part in several biochemical reactions:
| Enzyme | Reaction | Pathway |
| 2-hydroxyphytanoyl-CoA lyase | α-oxidation of phytanoic acid | |
| Alpha-ketoglutarate dehydrogenase complex | alpha-ketoglutarate -> succinyl-CoA | citric acid cycle |
| Branced-chain alpha-keto acid dehydrogenase complex | ||
| Pyruvate decarboxylase | pyruvate -> acetaldehyde | ethanol fermentation |
| Pyruvate dehydrogenase complex | pyruvate -> acetyl CoA | cellular respiration |
| Transketolase | pentose phosphate pathway |
TimeTree. Public knowledge-base for information on the tree-of-life and its evolutionary timescale. Three search modes are possible:
NODE TIME - to find the divergence time of two species or higher taxa
TIMELINE - to drill back through time and find evolutionary branches from the perspective of a single species
TIMETREE - to build a timetree of a group of species or custom list
Transcription. First step of gene expression, in which a part of DNA is copied into RNA.
The following steps of transcription can be recognized (Cramer 2019):
1. Promoter recognition
- Promoters often contain conserved DNA sequence elements, which differ for Pol I, Pol II and Pol III
- RNA polymerases cannot recognize promoter elements by themselves, they need transcription factors
- The RNA polymerases and transcription factors form specific pre-initiation complexes (PICs) on promoter DNA
2. Promoter opening
- Key function of the PIC is to open DNA
- In Pol I and Pol III systems, DNA is opened spontaneously with the use of binding energy alone
- DNA opening by Pol II generally requires an additional enzyme, the DNA translocase XPB, a subunit of the general factor TFIIH
- Dependency of Pol II on XPB means an additional layer of transcriptional regulation
3. Initiation regulation
- The formation, stability and function of the PIC are regulated in all three transcription systems
- Poll II initiation is regulated by the co-activator complex Mediator, that is composed of more than 20 proteins
- Mediator stimulates phosphorylation of the Pol II C-terminal domain (CTD) by the TFIIH kinase subunit CDK7 to facilitate the transition to the elongation phase of transcription
4. Elongation regulation
- An elongation complex forms when the RNA grows to a critical length
- To add a nucleotide to the growing RNA, the polymerase closes the active site, catalyses the formation of a phosphodiester bond using a two-metal-ion mechanism and moves to the next template position. Certain DNA sequences can, however, interrupt the nucleotide-addition cycle and induce transcriptional pausing
- Pausing can lead to polymerase backtracking, arrest and termination. Pol II can arrest in front of nucleosomes, but can be rescued by the elongation factor TFIIS. TFIIS binds the Pol II funnel and pore, aligns the DNA– RNA hybrid with the active site and triggers the cleavage of backtracked RNA to restart transcription
- In metazoan cells, the elongation phase of Pol II transcription is also regulated. Pol II often pauses about 50 base pairs downstream of the transcription start site, and such promoter-proximal pausing is highly regulated
- Promoter-proximal pausing can limit the frequency of transcription initiation, and thereby regulate a gene by changing the amount of RNA synthesized per unit of time
- The phosphorylated CTD recruits many factors that are required for Pol II elongation and for co-transcriptional events such as RNA processing, histone modification and chromatin remodelling
For transcription termination see (Proudfoot 2016)
Transcription factor (TF) protein capable of (1) binding DNA in a sequence-specific manner and (2) regulating transcription.
TFs form a complex system that guides expression of the genome. Over 1600 human TFs are known. Major classes of human TFs differ markedly in their evolutionary trajectories and expression patterns, underscoring distinct functions. TFs likewise underlie many different aspects of human physiology, disease, and variation, highlighting the importance of continued effort to understand TF-mediated gene regulation.
TFs have traditionally been classified as either "activators" or "repressors"; however this notion has been repeatedly questioned. Many TFs can recruit multiple cofactors that have opposite effects, dependent on the local sequence context and availability of cofactors.
Some major TF families in eukaryotes are C2H2-zinc finger (ZF), Homeodomain, basic helix-loop-helix (bHLH), basic leucine zipper (bZIP) and nuclear hormone receptor (NHR). (Lambert 2018).
Transcription Factory. a site containing at least two RNA polymerases (plus associated machinery) active on at least two different templates. The purpose behind this definition is to differentiate our use from two other cases. First, the term transcription “factory” has been applied to the various machines involved in the production of only one mature message (e.g., those involved in capping, splicing, and polyadenylation). Second, we wish to distinguish our factories from cases where two polymerases are active on the same template (Cook 2018).
Mid-1990's Cook et al made the observation that bromo-uridine triphosphate labeled transcripts in mammalian cells did not occur diffusely in the nucleplasm, but at discrete foci. Studies over the past twenty years have provided compelling evidence that Pol II and transcribing genes colocalize at specific foci in cells. It is still not clear however if the factories move to genes or genes move to factories (Buckley 2014)
Movie from Cook how an immobilized RNA polymerase might work: http://users.path.ox.ac.uk/~pcook/images/tcycle.html
Transposable element (TE, transposons, "jumping gene"). DNA sequence that has the ability to change position within a genome (Bourque 2018). TEs are major components of eukaryotic genomes and they have a yet unknown impact on genome evolution, function, and disease.
There are many forms and shapes of TEs, but they can be divided in two classes:
- Class I: retrotransposons. Mobilize through a "copy-and-paste" mechanism whereby a RNA intermediate is transcribed into a complementary DNA (cDNA) molecule, using the enzyme reverse transcriptase (RT). Different subclasses of retrotransposons use different mechanisms of chromosomal integration, using different types of enzymes. SINEs can only retrotranspose by using the RT and endonuclease activity of a partner LINE.
| Subclass | TE-encoded enzyme | Superfamily | Family |
| Long Terminal Repeat (LTR) | integrase | endogenous retroviruses (ERV) | Human endogenous retrovirus (HERV) |
| Non LTR | endonuclease | Long Interspersed Nuclear Element (LINE) | LINE-1 |
| Short Interspersed Nuclear Element (SINE) | Alu | ||
| Dictyostelium Repititive Sequence (DIRS) | tyrosine recombinase |
- Class II: DNA transposons. Transpose directly without an RNA intermediate.
TEs occupy a substantial portion of genomes. In maize, the species where Barbara McClintock (1983 Nobel Prize in Physiology or Medicine) discovered transposition (McClintock 1950), 60 to 70 % of the genome is comprised of LTR retrotransposons. More than 30 % of the human genome is comprised of LINEs and SINEs.
TEs are not randomly distributed in the genome. SINEs are generally localized in gene-rich regions, LINEs are enriched in intergenic regions (Elbarbary 2016).
Trans-regulatory element (TRE), opposite of Cis-regulatory element (CRE). Gene that regulates the expression of other genes, often situated on other molecules. TREs code voor transcription factors.
Tree of life. Tree-like representation of the relationship between organisms living on Earth. See TimeTree
Prior to the 1970s trees of life where mainly based on morphology and were confined to metazoa and metaphyta, whose histories at best cover 20% of the total evolutionary time span (Woese et al 1990). From the 1970s relations between organisms where increasingly determined by sequential information derived from DNA, RNA or proteins.
Based on the characteristics of small subunit ribosomal RNA (SSU rRNA) Woese et al (1977, 1990) proposed from the mid 1970s that the tree of life has three main branches (also called domains or superkingdoms): Archaea, Bacteria, and Eucarya.
More recent rearch methods and data suggest two hypotheses concerning the origin of the eukaryotic cell (Williams et al 2013):
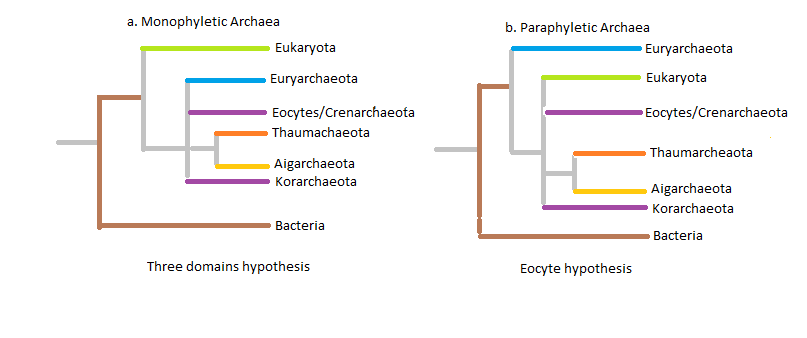
A modern version of the tree of life can be found in (Hug et al 2016): https://www.nature.com/articles/nmicrobiol201648
Tricarboxylic acid (TCA) cycle see Citric acid cycle
U - Z
Vertebrate Genomes Project (VGP). Rhie et al (2021).
Vitamins group of organic compounds which are essential for normal physiological functioning but which are not synthesised endogenously by the body and therefore have to be sequestered in small quantities from the diet. Although most vitamins are derived ultimately from plants, they are often consumed indirectly from higher up the food chain in foods of animal origin, including meat, dairy and eggs.
| Vitamin | Known as |
| B1 | Thiamine |
| B2 | Riboflavin |
| B3 | Niacin |
| B5 | Pantothenic acid |
| B6 | Pyridoxine |
| B7 | Biotin |
| B9 | Folate/ folic acid |
| B12 | cobalamins |
Zoonomia project is investigating the genomics of shared and specialized traits in eutherian mammals (Zoonomia Consortium 2020).
Literature
Literature
Aledo JC. 2019. Methionine in proteins: The Cinderella of the proteinogenic amino acids. Protein Science. 28:1785-1796. https://doi.org/10.1002/pro.3698
Benoist C, Mathis D. 2002. Mast cells in autoimmune disease. Nature. 420(6917):875-8. 10.1038/nature01324
Biesiekierski Jr. 2017. What is gluten?. Journal of Gasteroenterology and Hepatology. 32 (suppl 1): 78-81. https://doi.org/10.1111/jgh.13703
Bourque G et al. 2018. Ten things you should know about transposable elements. Genome Biology. 19: 199. https://doi.org/10.1186/s13059-018-1577-z
Bromilow S et al. 2017 Jun 23. A curated gluten protein sequence database to support development of proteomics methods for determination of gluten in gluten-free foods. J Proteomics. 163: 67–75. 10.1016/j.jprot.2017.03.026
Buckley SB and Lis JT; Imaging RNA Polymerase II transcription sites in living cells; Curr Opin Genet Dev 2014 April; 25: 126-130
Cook, P.R., and Marenduzzo, D. (2018). Transcription-driven genome organization: a model for chromosome structure and the regulation of gene expression tested through simulations. Nucl. Acids Res. 46, 9895-9906.
Cramer P; Organization and regulation of gene transcription; Nature 2019, 573: 45-54
De Las Rivas J et al; Protein–Protein Interactions Essentials: Key Concepts to Building and Analyzing Interactome Networks; PLoS Comput Biol. 2010 Jun; 6(6)
DeBernardinis RJ et al; Q's next: the diverse functions of glutamine in metabolism, cell biology and cancer; Oncogene (2010) 29: 313-324
Elbarbary RA et al; Retrotransposons as regulators of gene expression; Science 2016, 351: 679
ENCODE Project Consortium et al; Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007)
ENCODE Project Consortium et al; An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012)
ENCODE Project Consortium et al; Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020)
Froese DS et al. 2019. Vitamin B12, folate, and the methionine remethylation cycle - biochemistry, pathways, and regulation. J Inherit Metab Dis. 42:673-685
Goldstein B, King N; The Future of Cell Biology: Emerging Model Organisms; Trends Cell Biol. 2016 November, 26(11): 818-824
Goodwin S et al; Coming of age: ten years of next-generation sequencing technologies; Nat Rev Genet 17, 333–351 (2016)
Hakim O et al; Snapshot: Chromosome Conformation Capture; Cell. 148 (5): 1068.e1–2
Hug LA et al (11 April 2016). A new view of the tree of life. Nature Microbiology. 1 (5): 16048
Huxley-Jones J, Clarke TK, Beck C, Toubaris G, Robertson DL, Boot-Handford RP. 2007. The evolution of the vertebrate metzincins; insights from Ciona intestinalis and Danio rerio. BMC Evol Biol. 17;7:63. doi: 10.1186/1471-2148-7-63. PMID: 17439641; PMCID: PMC1867822.
Hnisz D et al. Super-enhancers in the control of cell identity and disease. Cell 2013 Nov 7;155(4):934-47
Kennedy DO; B Vitamins and the Brain: Mechanisms, Dose and Efficacy - A Review; Nutrients 2016, 8, 68
Koehl P; in Rev.Comput.Chem., K.B. Lipkowitz, T.R. Cundari, and V. Gillet, Eds, Wiley, New York, 2006, Vol. 22, pp 1-55. Protein Structure Classification. https://doi.org/10.1002/0471780367.ch1
Koonin EV; Orthologs, paralogs, and evolutionary genomics; Annual review of genetics 2005;39:309-38
Lambert, S.A. et al; The Human Transcription Factors; Cell 2018, 172: 650-665
Lenzen S; A Fresh View of Glycolysis and Glucokinase Regulation: History and Current Status; J Biol Chem 2014 May 2;289(18):12189-94. doi: 10.1074/jbc.R114.557314
Levi A; How evolution builds genes from scratch - Scientists long assumed that new genes appear when evolution tinkers with old ones. It turns out that natural selection is much more creative; Nature 17 october 2019, 574: 314–316
Luck K et al; A reference map of the human binary protein interactome. Nature 580, 402–408 (2020). https://doi.org/10.1038/s41586-020-2188-
McClintock, Barbara (1950) "The origin and behavior of mutable loci in maize". Proceedings of the National Academy of Sciences. 36:344–55
Metzker ML; Sequencing technologies - the next generation; Nature Reviews Genetics 11 (2010): 31-46
Mock DM; Biotin: From Nutrition to Therapeutics; J Nutr. 2017 Aug;147(8):1487-1492. doi: 10.3945/jn.116.238956
Meuleman W et al; Index and biological spectrum of human DNase I hypersensitive sites. Nature 584, 244–251 (2020). https://doi.org/10.1038/s41586-020-2559-3
Nathan C. 2002. Points of control in inflammation. Nature. 420: 846-52. https://doi.org/10.1038/nature01320
Nelson DL, Cox MM; Lehninger Principles of Biochemistry; sixth edition 2013
Osborne TB; The vegetable proteins; MONOGRAPHS ON BIOCHEMISTRY, LONGMANS, GREEN, AND CO 1909; The vegetable proteins
Owen OE et al; The Key Role of Anaplerosis and Cataplerosis for Citric Acid Cycle Function; The Journal of Biological Chemistry (2002) 277: 30409-30412
Pollard KS; An RNA gene expressed during cortical development evolved rapidly in humans; Nature. 2006 Sep 14;443(7108):167-72
Proudfoot NJ; Transcriptional termination in mammals: Stopping the RNA polymeraseII juggernaut; Science. 2016 Jun 10;352(6291)
Rhie A et al; Towards complete and error-free genome assemblies of all vertebrate species; Nature volume 592, pages737–746(2021)
Santos R et al; A comprehensive map of molecular drug targets; Nat Rev Drug Discov 16, 19–34 (2017). https://doi.org/10.1038/nrd.2016.230
Schübeler D; Function and information content of DNA methylation; Nature 2015, 517: 321-326
Scientific Advisory Committee on Nutrition; Carbohydrates and Health; 2015 https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/445503/SACN_Carbohydrates_and_Health.pdf
Shewry P; Cereal seed storage proteins: structures, properties and role in grain utilization; Journal of Experimental Botany (2002), Vol 53, No 370
Shewry P; What is gluten – why is it special?; Front Nutr. (2019) https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6625226/pdf/fnut-06-00101.pdf
Silhavy TJ et al; The Bacterial Cell Envelope; Cold Spring Harb Perspect Biol 2010;2:a000414
Vincze O et al; Cancer risk across mammals; Nature 2022,601: 263-267 https://doi.org/10.1038/s41586-021-04224-5
Williams TA et al; An archaeal origin of eukaryotes supports only two primary domains of life; Nature 2013, 504: 231-236
Woese CR, Fox GE. (1977) Phylogenetic structure of the prokaryotic domain: The primary kingdoms. Proc. Natl. Acad. Sci. USA 74, 5088-5090
Woese CR, Kandler O, Wheelis ML (1990). Towards a natural system of organisms: Proposal for the domains Archaea, Bacteria, and Eucarya. Proc. Natl. Acad. Sci. USA 87, 4576-4579
Xu D, Xu Y; Protein Databases on the Internet; Curr Protoc Mol Biol. 2004 Nov;Chapter 19:Unit 19.4. doi: 10.1002/0471142727.mb1904s68
Yang W; Nucleases: Diversity of Structure, Function and Mechanism; Q Rev Biophys . 2011 February ; 44(1): 1–93. doi:10.1017/S0033583510000181.
Zoonomia Consortium; A comparative genomics multitool for scientific discovery and conservation; Nature 2020, 587: 240-245; DOI: 10.1038/s41586-020-2876-6
.png)